Articles
- Page Path
- HOME > Endocrinol Metab > Volume 36(5); 2021 > Article
-
Original ArticleAdrenal Gland Metabolic Subtyping of Adrenal Tumors: Prospective Multi-Center Cohort Study in Korea
-
Eu Jeong Ku1*
 , Chaelin Lee2*
, Chaelin Lee2* , Jaeyoon Shim2, Sihoon Lee3, Kyoung-Ah Kim4, Sang Wan Kim5, Yumie Rhee6, Hyo-Jeong Kim7, Jung Soo Lim8, Choon Hee Chung8, Sung Wan Chun9, Soon-Jib Yoo10, Ohk-Hyun Ryu11, Ho Chan Cho12, A Ram Hong13, Chang Ho Ahn14, Jung Hee Kim15
, Jaeyoon Shim2, Sihoon Lee3, Kyoung-Ah Kim4, Sang Wan Kim5, Yumie Rhee6, Hyo-Jeong Kim7, Jung Soo Lim8, Choon Hee Chung8, Sung Wan Chun9, Soon-Jib Yoo10, Ohk-Hyun Ryu11, Ho Chan Cho12, A Ram Hong13, Chang Ho Ahn14, Jung Hee Kim15 , Man Ho Choi2
, Man Ho Choi2
-
Endocrinology and Metabolism 2021;36(5):1131-1141.
DOI: https://doi.org/10.3803/EnM.2021.1149
Published online: October 21, 2021
1Department of Internal Medicine, Chungbuk National University Hospital, Chungbuk National University College of Medicine, Cheongju, Korea
2Molecular Recognition Research Center, Korea Institute of Science and Technology, Seoul, Korea
3Department of Internal Medicine, Gachon University College of Medicine, Incheon, Korea
4Department of Internal Medicine, Dongguk University Ilsan Hospital, Dongguk University College of Medicine, Goyang, Korea
5Department of Internal Medicine, Seoul Metropolitan Government Seoul National University Boramae Medical Center, Seoul National University College of Medicine, Seoul, Korea
6Department of Internal Medicine, Yonsei University College of Medicine, Seoul, Korea
7Department of Internal Medicine, Nowon Eulji Medical Center, Eulji University, Seoul, Korea
8Department of Internal Medicine, Yonsei University Wonju College of Medicine, Wonju, Korea
9Department of Internal Medicine, Soonchunhyang University Cheonan Hospital, Soonchunhyang University College of Medicine, Cheonan, Korea
10Division of Endocrinology and Metabolism, Department of Internal Medicine, Bucheon St. Mary’s Hospital, College of Medicine, The Catholic University of Korea, Bucheon, Korea
11Department of Internal Medicine, Hallym University Chuncheon Sacred Heart Hospital, Hallym University College of Medicine, Chuncheon, Korea
12Department of Internal Medicine, Keimyung University School of Medicine, Daegu, Korea
13Department of Internal Medicine, Chonnam National University Medical School, Gwangju, Korea
14Department of Internal Medicine, Seoul National University Bundang Hospital, Seoul National University College of Medicine, Seongnam, Korea
15Department of Internal Medicine, Seoul National University Hospital, Seoul National University College of Medicine, Seoul, Korea
- Corresponding authors: Jung Hee Kim. Department of Internal Medicine, Seoul National University Hospital, Seoul National University College of Medicine, 101 Daehak-ro, Jongno-gu, Seoul 03080, Korea, Tel: +82-2-2072-4839, Fax: +82-2-762-9662, E-mail: jhee1@snu.ac.kr.
- Man Ho Choi. Molecular Recognition Research Center, Korea Institute of Science and Technology, 5 Hwarang-ro 14-gil, Seongbuk-gu, Seoul 02792, Korea, Tel: +82-2-958-5081, Fax: +82-2-958-5059, E-mail: mh_choi@kist.re.kr
- * These authors contributed equally to this work.
Copyright © 2021 Korean Endocrine Society
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (https://creativecommons.org/licenses/by-nc/4.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
ABSTRACT
-
Background
- Conventional diagnostic approaches for adrenal tumors require multi-step processes, including imaging studies and dynamic hormone tests. Therefore, this study aimed to discriminate adrenal tumors from a single blood sample based on the combination of liquid chromatography-mass spectrometry (LC-MS) and machine learning algorithms in serum profiling of adrenal steroids.
-
Methods
- The LC-MS-based steroid profiling was applied to serum samples obtained from patients with nonfunctioning adenoma (NFA, n=73), Cushing’s syndrome (CS, n=30), and primary aldosteronism (PA, n=40) in a prospective multicenter study of adrenal disease. The decision tree (DT), random forest (RF), and extreme gradient boost (XGBoost) were performed to categorize the subtypes of adrenal tumors.
-
Results
- The CS group showed higher serum levels of 11-deoxycortisol than the NFA group, and increased levels of tetrahydrocortisone (THE), 20α-dihydrocortisol, and 6β-hydroxycortisol were found in the PA group. However, the CS group showed lower levels of dehydroepiandrosterone (DHEA) and its sulfate derivative (DHEA-S) than both the NFA and PA groups. Patients with PA expressed higher serum 18-hydroxycortisol and DHEA but lower THE than NFA patients. The balanced accuracies of DT, RF, and XGBoost for classifying each type were 78%, 96%, and 97%, respectively. In receiver operating characteristics (ROC) analysis for CS, XGBoost, and RF showed a significantly greater diagnostic power than the DT. However, in ROC analysis for PA, only RF exhibited better diagnostic performance than DT.
-
Conclusion
- The combination of LC-MS-based steroid profiling with machine learning algorithms could be a promising one-step diagnostic approach for the classification of adrenal tumor subtypes.
- The prevalence of adrenal tumors increases with age, and ranges from 3% to 7% in adults over 50 years of age [1,2]. Most cases are incidentally detected with abdominal computed tomography or magnetic resonance imaging studies conducted for various irrelevant purposes [3]. According to recent guidelines, approximately 75% of all adrenal incidentalomas are nonfunctioning adenoma (NFA), and 15% are hormone-producing tumors, while adrenocortical carcinomas account for less than 10% [4]. However, the diagnostic approach for subtyping adrenal tumors requires multiple steps. For the detection of adrenal incidentalomas, 1 mg overnight dexamethasone suppression test (DST), 24-hour urinary fractionated metanephrine or plasma metanephrines, and plasma aldosterone to renin activity ratio for hormone excess secretion are necessary [5]. Even if the above-mentioned tests are performed, unresolved issues regarding indeterminate characteristics remain, and further tests are usually needed to confirm the diagnosis.
- In contrast to the cross-reactivity of conventional immunoassay techniques, the recent development of mass spectrometry has made highly selective profiling possible and is recommended for steroid analysis [6]. Understanding adrenal diseases has been accelerated by liquid chromatography-mass spectrometry (LC-MS)-based analyses [7–9]. Regarding adrenal diseases, LC-MS-based steroid profiling has characterized metabolic signatures of Cushing’s syndrome (CS), primary aldosteronism (PA), and congenital adrenal hyperplasia [10–12]. However, only few studies have simultaneously identified subtypes of adrenal tumors in a single run using LC-MS serum steroid profiling.
- Although LC-MS serum steroid profiling can provide relatively accurate measurements, to unleash the full potential of steroid profiles, the multidimensional diagnostic approach is required for recognizing and classifying patterns of steroids beyond traditional approaches to interpretation of results. Several steroids were not independent each other. Thus, the traditional statistical analyses have a limitation in controlling the interaction among several steroids. In this regard, other studies also applied the machine learning method in the steroid profiling analysis [13,14]. In conjunction with advances in bioanalytical technologies, machine learning algorithms have recently been introduced into medical diagnosis. Machine learning is a part of artificial intelligence data analysis that automates learning from big data, recognizing patterns, and providing diagnostic and predictive models [15,16]. Machine learning algorithms may also provide better diagnostic models for subtyping adrenal tumors as the performance of machine learning models is superior to that of traditional methods. In particular, if an algorithm is generated through the training process of classification and regression learning techniques from the LC-MS serum steroid profiles of a single sample, it might be possible to subtype adrenal tumors in response to input data (steroidomic data). Herein, we hypothesized that the combination of LC-MS-based steroid profiling with machine learning algorithms could streamline the categorization of adrenal tumors using a single blood sampling.
INTRODUCTION
- Study participants
- This study was performed as a part of an ongoing prospective multicenter study of adrenal diseases in South Korea, the Korean Adrenal Tumor Study (KATS). Consecutive patients who were diagnosed with adrenal diseases from June 2017 to October 2019 from 16 centers in Korea were enrolled. NFA, CS, and PA were diagnosed according to current guidelines [5,17]. CS was defined as an abnormal result (cut-off value for serum cortisol at 5.0 μg/dL) of the 1 mg overnight DST and/or as an increased 24-hour urinary free cortisol excretion in addition to clinical Cushing’s stigmata such as moon face, buffalo hump or red striae. Subjects (n=19) with mild autonomous cortisol secretion defined as a serum cortisol 1.9 to 5.0 μg/dL after 1 mg overnight DST were excluded from the analysis. PA was screened by aldosterone-to-renin ratio (ARR), adopting cutoff value of 30. Patients with a positive ARR were investigated one or more confirmatory tests by such as saline infusion or a high salt loading. PA was defined when aldosterone was not suppressed during confirmatory suppression testing (e.g., for the saline loading test, 10 ng/dL) [17]. In the final analysis, 143 patients aged 19 to 70 years were included. Written informed consent was obtained from all participants. The study protocol was approved by the Institutional Review Board (IRB) of Seoul National University Hospital (IRB No. 1704-060-845) and all participating institutions were approved by their Ethics Committee.
- Fasting serum samples for steroid profiling were collected between 6:00 and 8:00 AM and stored at −75°C. Medical records, including diabetes mellitus, hypertension, dyslipidemia, coronary heart disease, cerebrovascular disease, and menopausal status were noted. Height and body weight were measured during the enrollment. Participants were excluded for any of the following criteria: (1) pheochromocytoma/paraganglioma; (2) adrenocortical carcinoma; (3) within 5 years after diagnosis of malignancy; (4) major depressive disorder; (5) chronic alcoholism; (6) chronic renal impairment (creatinine clearance <30 mL/min); (7) liver cirrhosis; (8) current treatment with glucocorticoids, antihistamines, and contraceptives within three months before enrollment; and (9) other diseases and conditions that could cause an acute stress response, such as major surgery, acute coronary syndrome, and febrile disease within the 4 weeks prior to the enrollement.
- Serum steroid profiling
- Sample preparation was performed as described previously [9,16]. Briefly, Serum samples (200 μL) were spiked with 20 μL of an internal standard mixture (d4-cortsiol, d8-17α-hydroxyprogesterone, and d4-pregnenolone, 0.2 μg/mL; d9-progesterone and d3-17α-hydroxypregnenolone [17α-OHP], 0.1 μg/mL; d3-testosterone sulfate, 1 μg/mL; d3-testosterone, 0.02 μg/mL; d6-dehydroepiandrosterone [DHEA], 0.5 μg/mL). After dilution with 1.8 mL phosphate buffer (0.2 M, pH 7.2), the sample was incubated with 50 μL of β-glucuronidase extracted from Escherichia coli (aqueous solution stabilized with 50% glycerol; Roche Diagnostics GmbH, Mannheim, Germany) for 1 hour at 55°C. The hydrolyzed sample was loaded on an Oasis HLB cartridge (3 mL, 60 mg; Waters, Milford, MA, USA) preconditioned with 4 mL of methanol, followed by 4 mL of distilled water (Burdick & Jackson, Muskegon, MI, USA) for solid-phase extraction. The cartridge was washed twice with 10% methanol (0.7 mL). Serum adrenal steroids were then eluted with absolute methanol (1 mL×2). The combined eluates were evaporated under a stream of nitrogen at 40°C. The dried extract was reconstituted with 50 μL of methanol and centrifuged using an Ultrafree-MC Centrifugal Filter (polyvinylidene fluoride, pore size: 0.1 μm; Millipore, Billerica, MA, USA) for 5 minutes at 14,000 rpm. Subsequently, 50 μL of 10% dimethyl sulfoxide was added to the Ultrafree-MC filter and centrifuged for 5 minutes at 14,000 rpm. Finally, an aliquot (5 μL) was injected into the LC-MS system.
- Statistical analysis
- Continuous variables were expressed as mean±standard deviation and analyzed using analysis of variance (ANOVA). Categorical variables are presented as numbers with percentages. Categorical variables among groups were compared using the chi-square test with Fisher’s exact test. Serum steroid levels among groups are presented as medians with interquartile ranges due to skewed data. The log-transformed serum steroid panels were compared among groups using ANOVA and analysis of covariance (ANCOVA) adjusted for age and sex. Multiple testing correction was conducted using the post hoc analysis with Tukey’s honestly significant difference test. Statistical analyses were conducted using SPSS version 20.0 for Windows (IBM Co., Armonk, NY, USA) and GraphPad Prism version 4.0 (GraphPad, La Jolla, CA, USA). In this study, a P value less than 0.05 was considered statistically significant.
- Machine learning analyses were performed to identify featured steroids and elucidate diagnostic models for adrenal tumors. The quantitative results of steroid profiling were processed by centering and scaling. We randomly divided the training (n=115) and test dataset (n=28) at a ratio of 8:2 and conducted 10 times, 5-fold cross-validation to reduce the overfitting of variables in the training dataset. Subsequently, three machine learning methods for classification with a decision tree, random forest, and extreme gradient boost (XGBoost) were applied. The decision tree model was formed by a multilevel split based on the serum steroids with high sensitivity and specificity in the receiver operating characteristic (ROC) curve analysis. Each node corresponds to the best cutoff threshold of the serum steroids for discriminating NFA, CS, and PA. Random forest is a combination of decision tree predictors where each tree depends on independently sampled random vector values and has the same distribution for all trees in the forest [18]. XGBoost is a decision-tree-based gradient boosting algorithm that avoids overfitting and bias [19]. The area under the curve for the receiver operating characteristic (AUROC) curve of each model was calculated to evaluate the diagnostic power of each decision tree, random forest, and XGBoost algorithm. The AUROC curves derived from each method for diagnosing CS and PA were compared using DeLong’s method. Machine learning analyses were performed using R version 3.6.2 (Foundation for Statistical Computing, Vienna, Austria). Various R packages, namely, tree, catools, caret, and XGBoost were used.
METHODS
- Clinical and laboratory characteristics of study subjects
- The clinical characteristics and hormonal values of patients with NFA, CS, and PA are listed in Table 1. The mean age of the NFA group was higher than that of the CS and PA groups. The percentage of women (24/30, 80.0%) was highest in the CS group (P=0.006), but the proportion of menopausal women was highest in the NFA group (P=0.004). The PA group showed the highest prevalence of hypertension (39/40, 97.5%) among the three groups (P<0.001). The BMI was similar among all the groups. There were no significant differences in comorbidities such as diabetes mellitus, dyslipidemia, coronary heart disease, and cerebrovascular disease among the groups.
- Comparison of serum steroid profiles
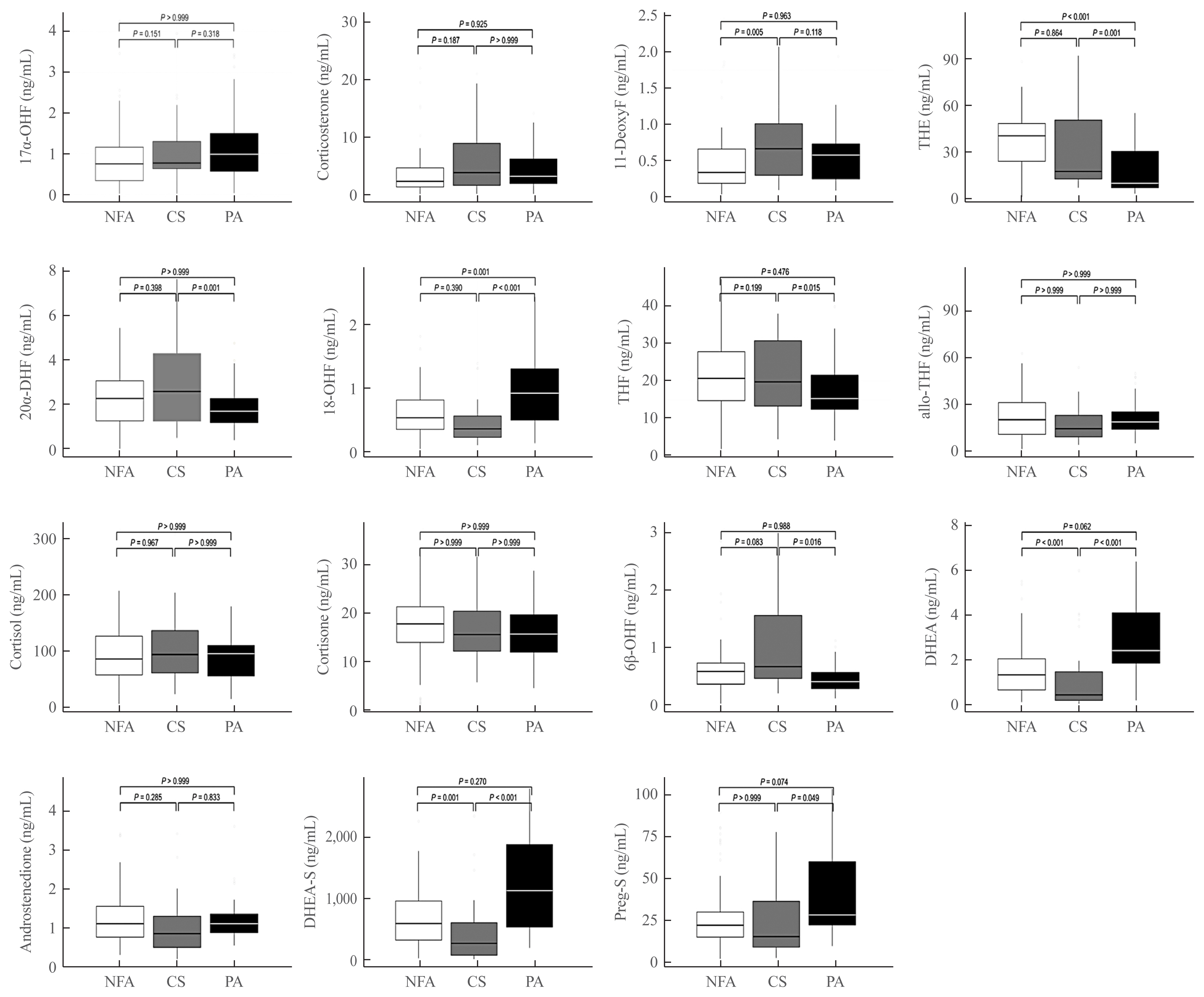
- Serum steroid profiles adjusted for age and sex among the groups were compared between the groups studied (Fig. 1). Compared to the NFA group, the CS group showed higher levels of 11-deoxycortisol (11-deoxyF), whereas both DHEA and DHEA-sulfate (DHEA-S) levels were found to have decreased in this group (P=0.005, P<0.001, and P=0.001, respectively). The PA group showed significantly decreased levels of tetrahydrocortisone (THE) and increased levels of 18-hydroxycortisol (18-OHF) as compared to the NFA group (P=0.001 amd P=0.001, respectively). When comparing the CS and PA groups, higher levels of THE, 20α-dihydrocortisol (20α-DHF), tetrahydrocortisol (THF), and 6β-hydroxycortisol (6β-OHF) were detected in the CS group, whereas higher levels of 18-OHF, DHEA, and DHEA-S were observed in the PA group (P=0.001, P=0.001, P=0.015, P=0.016, P<0.001, P<0.001, and P<0.001, respectively).
- Machine learning-based diagnostic models for subtyping adrenal tumors
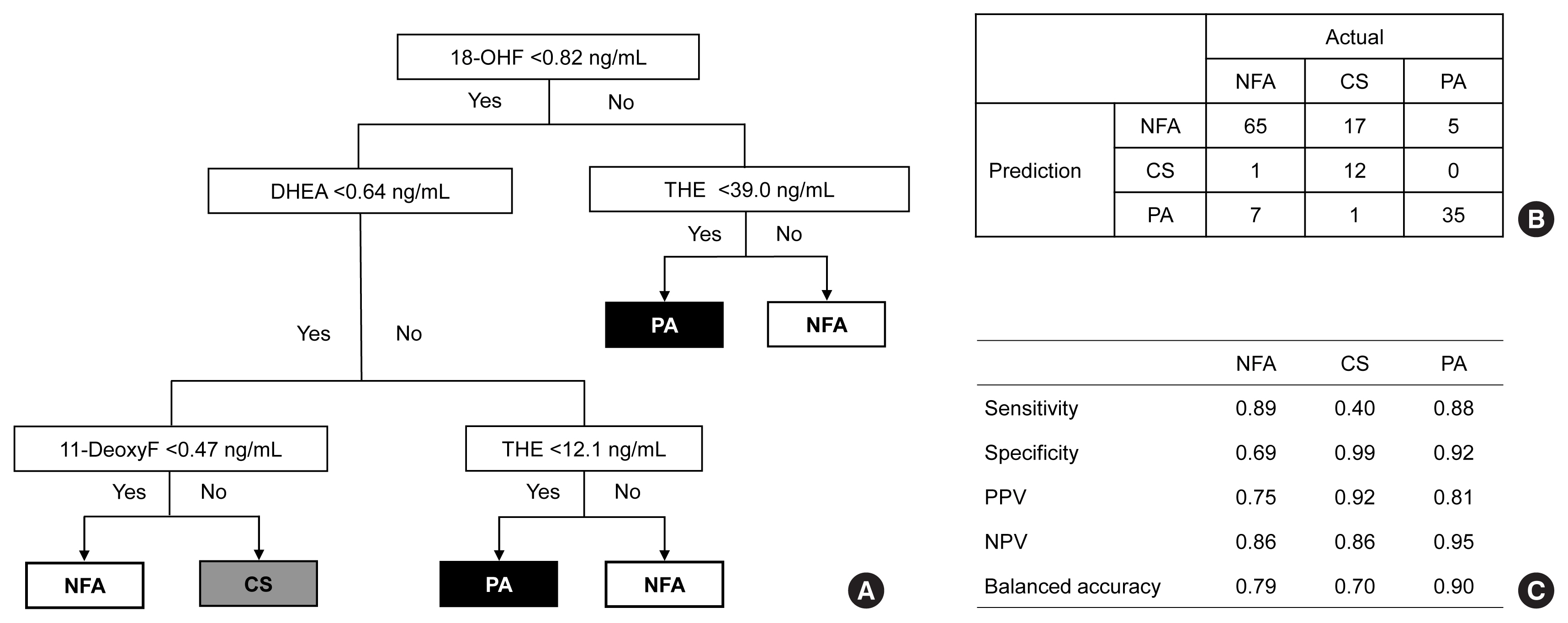
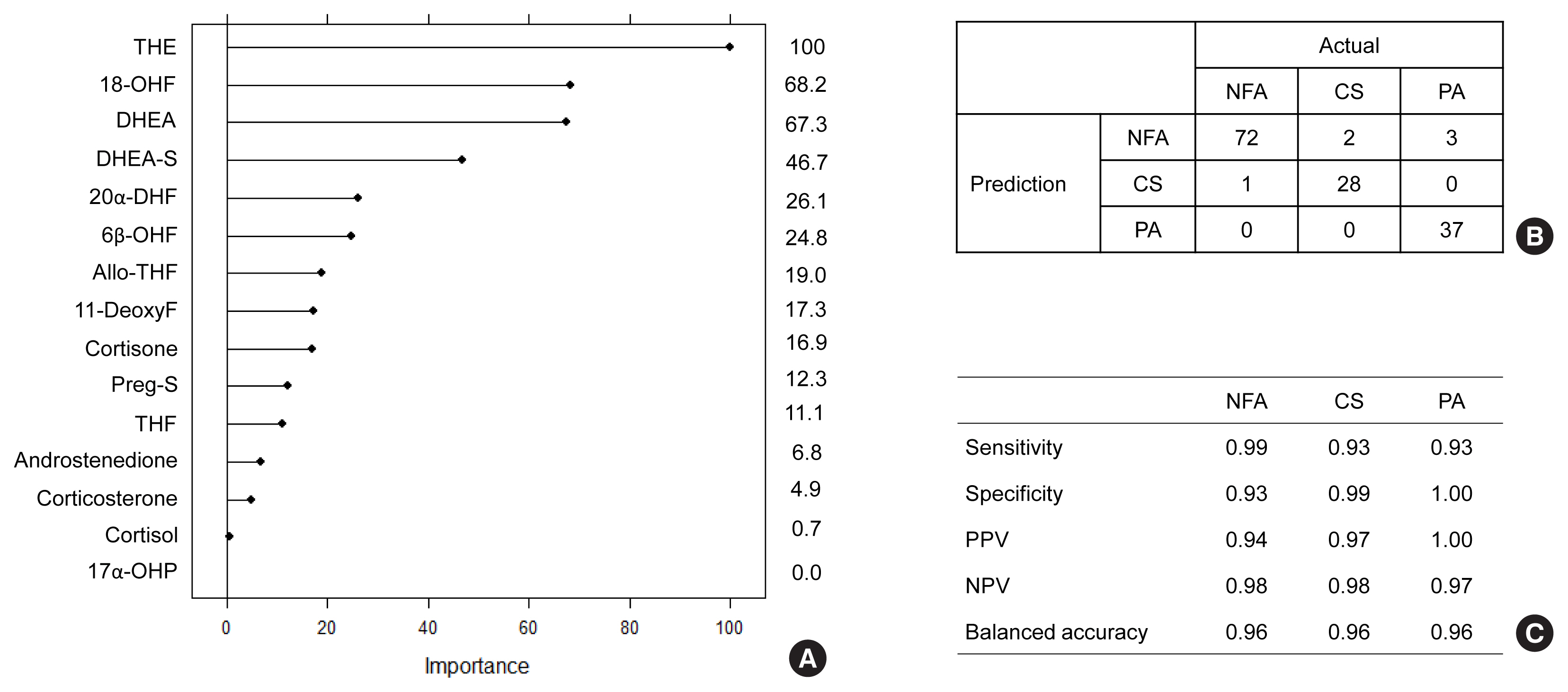
- Decision tree analysis was first applied to classify the NFA, CS, and PA groups in subjects with adrenal tumors using multiple-steroid panels (Fig. 2). Among the 15 steroid panels, 18-OHF, DHEA, THE, and 11-deoxyF were used as tree predictors (Fig. 2A). In the confusion matrix, decision tree analysis was classified correctly into each group at the rate of 78% (95% confidence interval [CI], 71% to 85%; P<1.6×10−11) (Fig. 2B). The balanced accuracy was found to be the lowest in the CS group, with the lowest sensitivity and highest specificity among the three groups. In the PA group, the balanced accuracy was 90%, which was the highest among the three groups (Fig. 2C).
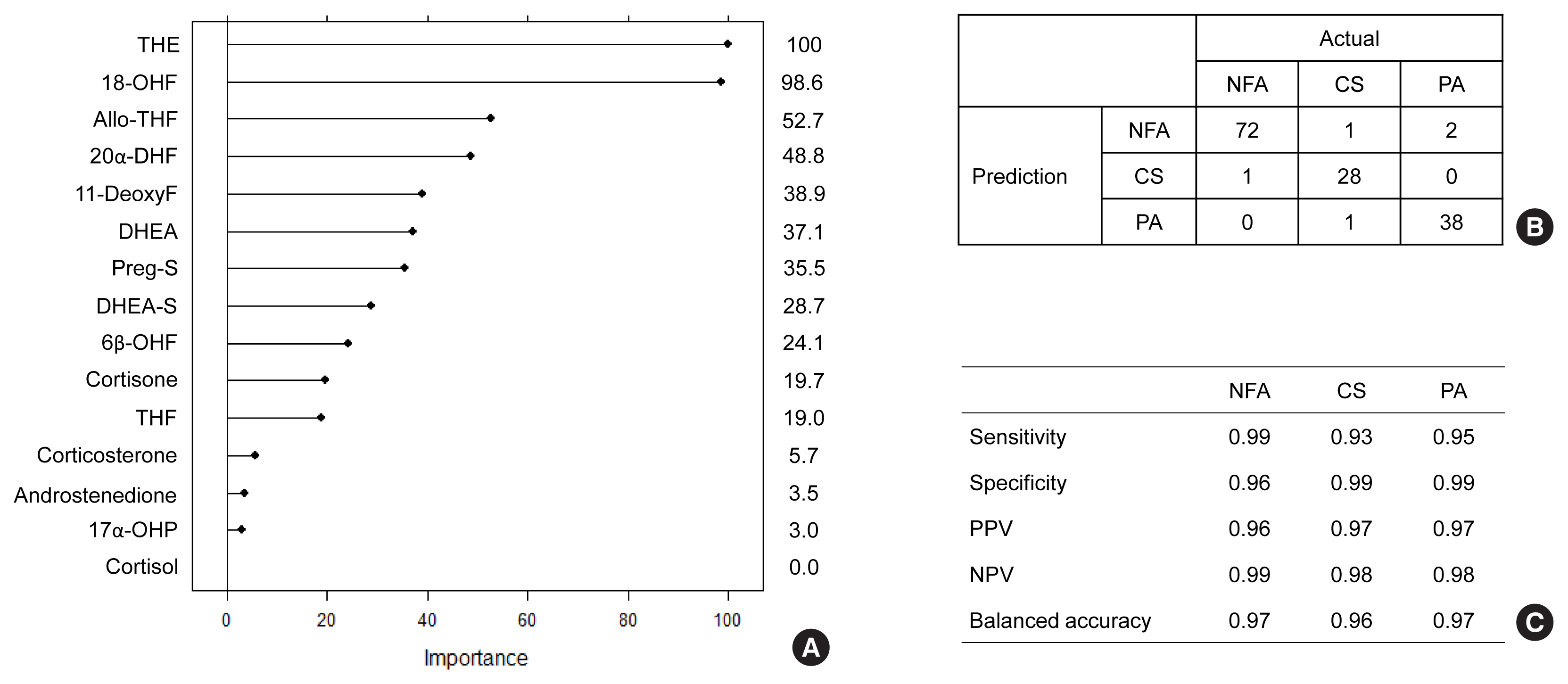
- Second, we used a random forest algorithm (Fig. 3). Based on their importance, the top three steroids were THE, 18-OHF, and DHEA (Fig. 3A). In the confusion matrix, random forest was classified correctly into each group at the rate of 96% (95% CI, 91% to 98%; P<2×10−16) (Fig. 3B). All three group exhibited similar balanced accuracy of 96% (Fig. 3C).
- The application of the XGBoost algorithm improved the overall predictive performance for discriminating adrenal tumors (Fig. 4). Among the 15 steroids featured, THE, and 18-OHF were found to be of high importance (Fig. 4A). The confusion matrix of XGBoost presented an accuracy of 97% (95% CI, 92% to 99%; P<2.0×10−16) (Fig. 4B). It accurately classified NFA, CS, and PA with balanced accuracies of 97%, 96%, and 97%, respectively (Fig. 4C).
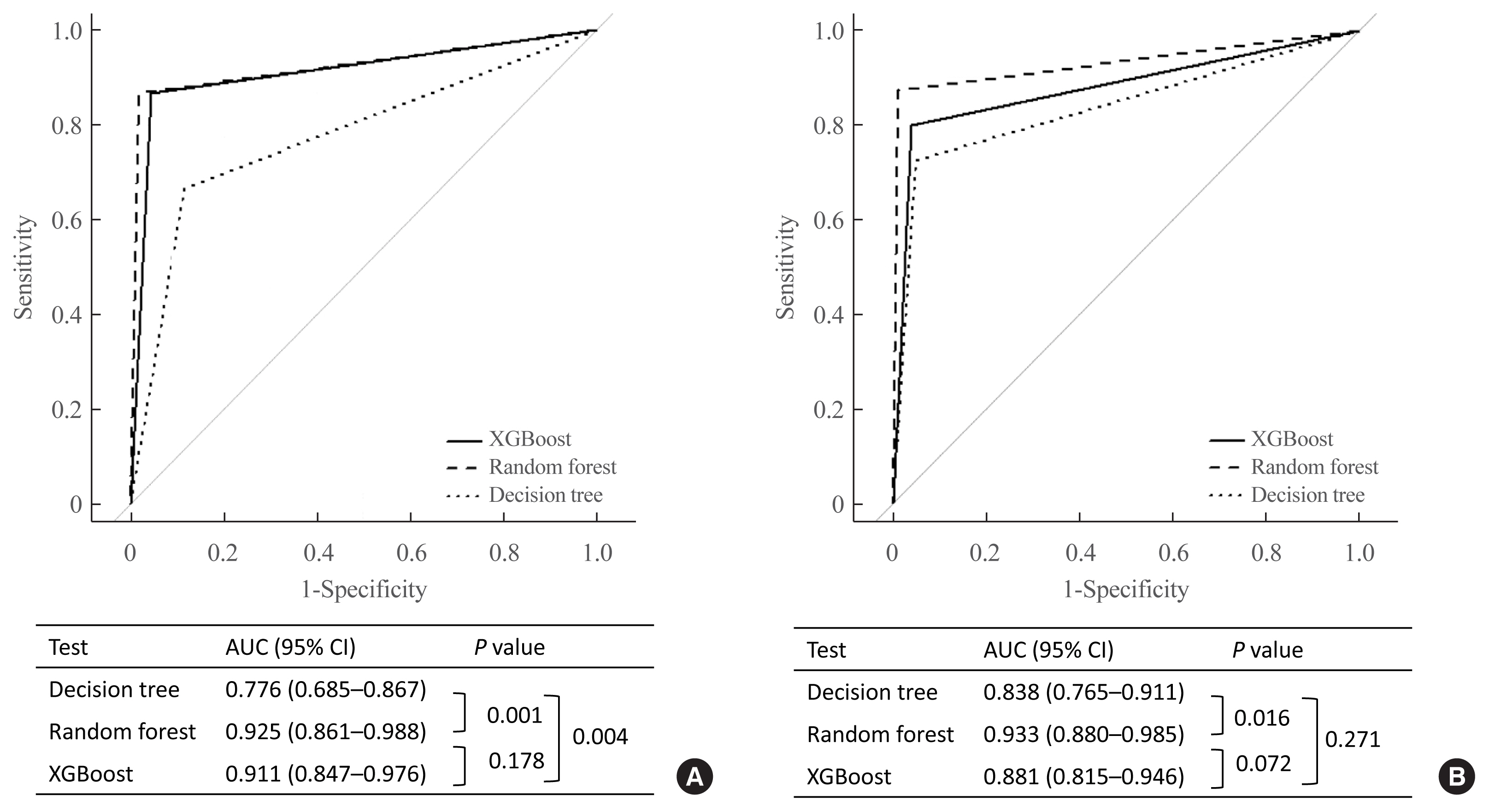
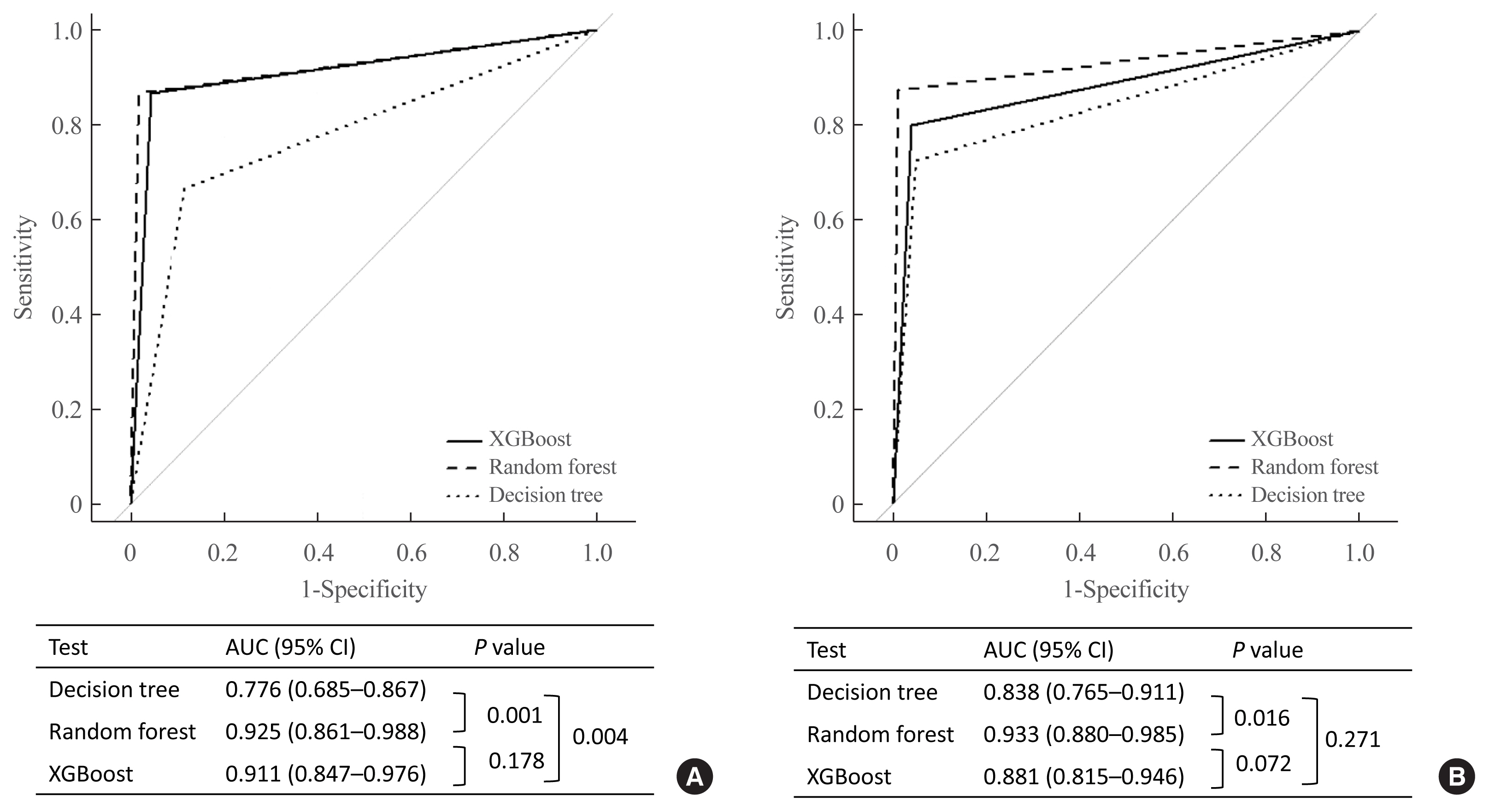
- In the ROC curve analysis, we validated the discrimination power of the decision tree analysis, random forest, and XGBoost algorithms for each subtype. For diagnosing CS, the area under the curves (AUCs) of the XGBoost algorithm and the random forest were significantly higher than those of the decision tree analysis (AUC, 0.911 [95% CI, 0.847 to 0.976], 0.925 [95% CI, 0.861 to 0.988], and 0.776 [95% CI, 0.685 to 0.867], respectively) (Fig. 5A). However, for PA, the AUC of the random forest was highest among the three methods (AUC, 0.933 [95% CI, 0.880 to 0.985]) (Fig. 5B).
RESULTS
- In this study, LC-MS-based simultaneous analysis of 15 adrenal steroids in serum samples obtained from patients with adrenal tumors was performed. In particular, we highlighted the following three discriminatory steroids for NFA, CS, and PA using the decision trees, random forest, and XGBoost analysis: THE, 18-OHF, and DHEA. The random forest algorithm of steroid profiling showed the highest diagnostic power for identifying CS and PA from single blood sampling.
- Subjects with CS exhibited a higher level of serum 11-deoxyF than those with NFA, while higher levels of serum THE, THF, 20α-DHF, and 6β-OHF than those in PA were observed (Fig. 1, Supplemental Table S1). These findings may implicate an increase in overall glucocorticoid precursors and metabolites in subjects with CS, which is in accordance with a recent study regarding the diagnosis of CS [20]. In a retrospective cross-sectional study measuring 15 plasma steroids, the CS group showed elevated levels of a cortisol precursor of 11-deoxyF compared with the control group [21]. Previous studies comparing the CS group with the control or NFA group reported increased levels of 11-deoxyF in the CS group [22–24]. The increase in 11-deoxyF was not related to the 17α-OHP level but instead was related to cortisol level. This finding may suggest hyperactivation of 11β-hydroxylase, which converts 11-deoxyF to cortisol, rather than its dysregulation. However, the levels of other glucocorticoid precursors and metabolites, such as THE, 20α-DHF, THF, and 6β-OHF did not differ between the CS and NFA groups.
- Other distinctive steroid features of the CS group were low DHEA and DHEA-S levels. DHEA is sulfated by DHEA sulfotransferase 2A1 to form DHEA-S [25]. Both DHEA and DHEA-S adrenal androgens are secreted by the zona reticularis of the adrenal gland under the dominant control of adrenocorticotropic hormone (ACTH). The suppression of ACTH driven by cortisol excess in adrenal CS reduces androgen production in the adjacent normal adrenal cortex. Both adrenal steroids have been suggested as valuable markers for differenting adrenal CS from pituitary CS [16,21,26]. Since the degree of hypercortisolemia and ACTH inhibition are proportional, previous studies have demonstrated that DHEA and DHEA-S levels could distinguish overt CS from subclinical CS and subclinical CS from NFA [8,22,24,27]. Sine DHEA-S was more abundant than DHEA, commercial immunoassays for DHEA-S were available. However, DHEA-S measurement is not approved as the standard diagnostic test for CS because the reference range varies according to age, sex, and assay [28–30]. Our study suggests that suppressed DHEA and DHEA-S, combined with other steroid profiling, can be used as significant variables for identifying CS.
- In the current study, the distinctive steroid in the PA group was 18-OHF. Being one of the adrenal cortical hybrid steroids, 18-OHF, has negligible glucocorticoid activity and no mineralocorticoid activity [30,31]. However, increased excretion of 18-OHF was identified in the urine, serum, and specifically adrenal venous samples of patients with PA [32,33]. Moreover, a high level of 18-OHF was associated with the potassium inwardly rectifying channel subfamily J member 5 (KCNJ5) somatic mutation in aldosterone-producing adenomas [34]. Given the high prevalence of KCNJ5 somatic mutation in Asian aldosterone-producing adenoma patients, 18-OHF can be used as an additional biomarker for subtyping PA [35].
- As the next step, the machine learning algorithm, i.e., random forest and XGBoost, was conducted to subtype adrenal tumors using multi-collinear steroid panels simultaneously. THE was the most featured steroid among the 15 steroids, and its levels in the CS and NFA groups were higher than in the PA; however, no difference was observed between the CS and NFA groups. While cortisol is metabolized to THF or allo-THF, cortisone, which is interconverted from cortisol by the 11β-hydroxysteroid dehydrogenase system, is metabolized to THE [25]. The tetrahydro metabolites of glucocorticoids account for more than 50% of urinary glucocorticoid metabolites. Since excessive cortisol is inactivated to cortisone by 11β-hydroxysteroid dehydrogenase type 2 in the CS patients, the cortisone metabolite, THE, can be dominantly elevated.
- This prospective multicenter cohort study for adrenal tumors was conducted using a unified protocol for sample preparation and transport according to the standard operating procedure. Moreover, the steroid profiling analysis was conducted at a single experienced core center. The final analysis was performed using a machine learning algorithm, and the present study was the first to discriminate the three different adrenal tumors simultaneously with the single-step serum steroid profiling.
- Despite these strengths, this study also has several limitations. First, steroid profiling of the patient groups was not compared with that of healthy subjects. Second, machine learning analysis with a small sample size may overfit the model [36]. To avoid overfitting bias, we split the training and test sets and applied 10-times, 5-fold cross-validation. However, the diagnostic model was not externally validated in another independent study group [37]. Third, due to the lack of sample size, patients with mild autonomous cortisol secretion were not included. Therefore, there are necessities to further analyze the mild autonomous cortisol secretion group as an independent group with a sufficient sample size in the future study. Fourth, the study included only Korean patients. Hence, the specific cut-offs of multiple steroid profiles for adrenal tumor subtypes could not be generalized for all races.
- In conclusion, quantitative steroid profiling could help discriminate between NFA, CS, and PA with promising accuracy. The identified differences in steroid profiles have significantly improved sensitivity and specificity after applying machine learning algorithms. This study demonstrates high diagnostic performance for the discrimination potency of steroid markers selected by mass spectrometry-based steroid profiling and machine learning algorithms in a single blood sampling. This promising one-step diagnostic approach for classifying adrenal tumor subtypes should be further validated in a large-scale prospective cohort study.
DISCUSSION
-
Acknowledgements
- Some of the biospecimens and data used in this study were provided by the Biobank of Seoul National University Hospital, a member of the Korea Biobank Network.
- This study was supported by the EnM research award from the Korean Endocrinology Society in 2017. Also, this study was conducted with the support of the Korea Institute of Science and Technology Institutional Program (Project No. 2E31093), the National Research Foundation by the Ministry of Science and ICT of Korea (Project No. NRF-2020R1C1C1010723), and a grant of the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health and Welfare of the Republic of Korea (Project No. HI21C0032). The funding source had no role in the design, data collection, analysis, interpretation of data, writing of the manuscript, or in the decision to submit the manuscript for publication. All authors contributed to this study, commented on the manuscript, and agreed to submit the final version of the manuscript for publication.
SUPPLEMENTARY INFORMATION
Supplemental Table S1.
-
CONFLICTS OF INTEREST
No potential conflict of interest relevant to this article was reported.
-
AUTHOR CONTRIBUTIONS
Conception or design: J.H.K., M.H.C. Acquisition, analysis, or interpretation of data: E.J.K., C.L., J.S., S.L., K.A.K., S.W.K., Y.R., H.J.K., J.S.L., C.H.C., S.W.C., S.J.Y., O.H.R., H.C.C., A.R.H., C.H.A., J.H.K., M.H.C. Drafting the work or revising: E.J.K., J.H.K., M.H.C. Final approval of the manuscript: E.J.K., C.L., J.S., S.L., K.A.K., S.W.K., Y.R., H.J.K., J.S.L., C.H.C., S.W.C., S.J.Y., O.H.R., H.C.C., A.R.H., C.H.A., J.H.K., M.H.C.
Article information

Values are expressed as mean±standard deviation, number (%), or median (interquartile range). The P values were calculated using Kruskal-Wallis or a one-way analysis of variance (ANOVA). The proportion of postmenopausal women was expressed as the proportion for women in each group.
NFA, nonfunctioning adenoma; CS, Cushing’s syndrome; PA, primary aldosteronism; BMI, body mass index; HTN, hypertension; DM, diabetes mellitus; CHD, coronary heart disease; CVA, cerebrovascular accident; DST, dexamethasone suppression test; UFC, urinary free cortisol; ACTH, adrenocorticotropic hormone; PRA, plasma renin activity; ARR, aldosterone-to-renin ratio.
- 1. Kloos RT, Gross MD, Francis IR, Korobkin M, Shapiro B. Incidentally discovered adrenal masses. Endocr Rev 1995;16:460–84.ArticlePubMed
- 2. Grossman A, Koren R, Tirosh A, Michowiz R, Shohat Z, Rahamimov R, et al. Prevalence and clinical characteristics of adrenal incidentalomas in potential kidney donors. Endocr Res 2016;41:98–102.ArticlePubMed
- 3. Young WF Jr. Clinical practice. The incidentally discovered adrenal mass. N Engl J Med 2007;356:601–10.ArticlePubMed
- 4. Fassnacht M, Arlt W, Bancos I, Dralle H, Newell-Price J, Sahdev A, et al. Management of adrenal incidentalomas: European Society of Endocrinology Clinical Practice Guideline in collaboration with the European Network for the Study of Adrenal Tumors. Eur J Endocrinol 2016;175:G1–34.ArticlePubMed
- 5. Lee JM, Kim MK, Ko SH, Koh JM, Kim BY, Kim SW, et al. Clinical guidelines for the management of adrenal incidentaloma. Endocrinol Metab (Seoul) 2017;32:200–18.ArticlePubMedPMC
- 6. Choi MH. Mass spectrometry-based steroid profiling: meeting unmet clinical needs. Tohoku J Exp Med 2021;253:171–80.ArticlePubMed
- 7. Arlt W, Biehl M, Taylor AE, Hahner S, Libe R, Hughes BA, et al. Urine steroid metabolomics as a biomarker tool for detecting malignancy in adrenal tumors. J Clin Endocrinol Metab 2011;96:3775–84.ArticlePubMedPMC
- 8. Di Dalmazi G, Fanelli F, Mezzullo M, Casadio E, Rinaldi E, Garelli S, et al. Steroid profiling by LC-MS/MS in nonsecreting and subclinical cortisol-secreting adrenocortical adenomas. J Clin Endocrinol Metab 2015;100:3529–38.ArticlePubMed
- 9. Lee C, Kim JH, Moon SJ, Shim J, Kim HI, Choi MH. Selective LC-MRM/SIM-MS based profiling of adrenal steroids reveals metabolic signatures of 17α-hydroxylase deficiency. J Steroid Biochem Mol Biol 2020;198:105615.ArticlePubMed
- 10. Loriaux DL. Diagnosis and differential diagnosis of Cushing’s syndrome. N Engl J Med 2017;376:1451–9.ArticlePubMed
- 11. Peitzsch M, Dekkers T, Haase M, Sweep FC, Quack I, Antoch G, et al. An LC-MS/MS method for steroid profiling during adrenal venous sampling for investigation of primary aldosteronism. J Steroid Biochem Mol Biol 2015;145:75–84.ArticlePubMed
- 12. El-Maouche D, Arlt W, Merke DP. Congenital adrenal hyperplasia. Lancet 2017;390:2194–210.ArticlePubMed
- 13. Eisenhofer G, Duran C, Cannistraci CV, Peitzsch M, Williams TA, Riester A, et al. Use of steroid profiling combined with machine learning for identification and subtype classification in primary aldosteronism. JAMA Netw Open 2020;3:e2016209.ArticlePubMedPMC
- 14. Chortis V, Bancos I, Nijman T, Gilligan LC, Taylor AE, Ronchi CL, et al. Urine steroid metabolomics as a novel tool for detection of recurrent adrenocortical carcinoma. J Clin Endocrinol Metab 2020;105:e307–18.ArticlePubMed
- 15. Erlic Z, Reel P, Reel S, Amar L, Pecori A, Larsen CK, et al. Targeted metabolomics as a tool in discriminating endocrine from primary hypertension. J Clin Endocrinol Metab 2021;106:1111–28.ArticlePubMed
- 16. Ahn CH, Lee C, Shim J, Kong SH, Kim SJ, Kim YH, et al. Metabolic changes in serum steroids for diagnosing and subtyping Cushing’s syndrome. J Steroid Biochem Mol Biol 2021;210:105856.ArticlePubMed
- 17. Funder JW, Carey RM, Mantero F, Murad MH, Reincke M, Shibata H, et al. The management of primary aldosteronism: case detection, diagnosis, and treatment: an Endocrine Society Clinical Practice Guideline. J Clin Endocrinol Metab 2016;101:1889–916.ArticlePubMed
- 18. Breiman L. Random forests. Mach Learn 2001;45:5–32.
- 19. Chen T, Guestrin C. XGBoost: a scalable tree boosting system. arXiv 2016 Jun 10 https://arxiv.org/abs/1603.02754 .
- 20. Nieman LK, Biller BM, Findling JW, Newell-Price J, Savage MO, Stewart PM, et al. The diagnosis of Cushing’s syndrome: an Endocrine Society Clinical Practice Guideline. J Clin Endocrinol Metab 2008;93:1526–40.ArticlePubMedPMC
- 21. Eisenhofer G, Masjkur J, Peitzsch M, Di Dalmazi G, Bidlingmaier M, Gruber M, et al. Plasma steroid metabolome profiling for diagnosis and subtyping patients with Cushing syndrome. Clin Chem 2018;64:586–96.ArticlePubMed
- 22. Masjkur J, Gruber M, Peitzsch M, Kaden D, Di Dalmazi G, Bidlingmaier M, et al. Plasma steroid profiles in subclinical compared with overt adrenal Cushing syndrome. J Clin Endocrinol Metab 2019;104:4331–40.ArticlePubMed
- 23. Katz FH, Lipman MM, Frantz AG, Jailer JW. The physiologic significance of 6beta-hydroxycortisol in human corticoid metabolism. J Clin Endocrinol Metab 1962;22:71–7.PubMed
- 24. Di Dalmazi G, Fanelli F, Zavatta G, Ricci Bitti S, Mezzullo M, Repaci A, et al. The steroid profile of adrenal incidentalomas: subtyping subjects with high cardiovascular risk. J Clin Endocrinol Metab 2019;104:5519–28.ArticlePubMed
- 25. Melmed S, Williams RH. Williams textbook of endocrinology; 14th ed. Philadelphia: Elsevier; 2019. Chapter 15:The adrenal cortex. p. 480–541.
- 26. Yener S, Yilmaz H, Demir T, Secil M, Comlekci A. DHEAS for the prediction of subclinical Cushing’s syndrome: perplexing or advantageous? Endocrine 2015;48:669–76.ArticlePubMed
- 27. Cunningham SK, McKenna TJ. Dissociation of adrenal androgen and cortisol secretion in Cushing’s syndrome. Clin Endocrinol (Oxf) 1994;41:795–800.ArticlePubMed
- 28. Pepping J. DHEA: dehydroepiandrosterone. Am J Health Syst Pharm 2000;57:2048–50.ArticlePubMed
- 29. Perrini S, Laviola L, Natalicchio A, Giorgino F. Associated hormonal declines in aging: DHEAS. J Endocrinol Invest 2005;28(3 Suppl):85–93.
- 30. Ulick S, Chu MD. Hypersecretion of a new corticosteroid, 18-hydroxycortisol in two types of adrenocortical hypertension. Clin Exp Hypertens A 1982;4:1771–7.ArticlePubMed
- 31. Lenders J, Williams TA, Reincke M, Gomez-Sanchez CE. Diagnosis of endocrine disease: 18-oxocortisol and 18-hydroxycortisol. Is there clinical utility of these steroids? Eur J Endocrinol 2018;178:R1–9.ArticlePubMed
- 32. Mulatero P, di Cella SM, Monticone S, Schiavone D, Manzo M, Mengozzi G, et al. 18-Hydroxycorticosterone, 18-hydroxycortisol, and 18-oxocortisol in the diagnosis of primary aldosteronism and its subtypes. J Clin Endocrinol Metab 2012;97:881–9.ArticlePubMed
- 33. Eisenhofer G, Dekkers T, Peitzsch M, Dietz AS, Bidlingmaier M, Treitl M, et al. Mass spectrometry-based adrenal and peripheral venous steroid profiling for subtyping primary aldosteronism. Clin Chem 2016;62:514–24.ArticlePubMed
- 34. Hong AR, Kim JH, Song YS, Lee KE, Seo SH, Seong MW, et al. Genetics of aldosterone-producing adenoma in Korean patients. PLoS One 2016;11:e0147590.ArticlePubMedPMC
- 35. Satoh F, Morimoto R, Ono Y, Iwakura Y, Omata K, Kudo M, et al. Measurement of peripheral plasma 18-oxocortisol can discriminate unilateral adenoma from bilateral diseases in patients with primary aldosteronism. Hypertension 2015;65:1096–102.ArticlePubMed
- 36. Hawkins DM. The problem of overfitting. J Chem Inf Comput Sci 2004;44:1–12.ArticlePubMed
- 37. Butler KT, Davies DW, Cartwright H, Isayev O, Walsh A. Machine learning for molecular and materials science. Nature 2018;559:547–55.ArticlePubMed
References
Figure & Data
References
Citations

- Treating Primary Aldosteronism-Induced Hypertension: Novel Approaches and Future Outlooks
Nathan Mullen, James Curneen, Padraig T Donlon, Punit Prakash, Irina Bancos, Mark Gurnell, Michael C Dennedy
Endocrine Reviews.2024; 45(1): 125. CrossRef - Steroid profiling in adrenal disease
Danni Mu, Dandan Sun, Xia Qian, Xiaoli Ma, Ling Qiu, Xinqi Cheng, Songlin Yu
Clinica Chimica Acta.2024; 553: 117749. CrossRef - Serum and hair steroid profiles in patients with nonfunctioning pituitary adenoma undergoing surgery: A prospective observational study
Seung Shin Park, Yong Hwy Kim, Ho Kang, Chang Ho Ahn, Dong Jun Byun, Man Ho Choi, Jung Hee Kim
The Journal of Steroid Biochemistry and Molecular Biology.2023; 230: 106276. CrossRef - Recent Updates on the Management of Adrenal Incidentalomas
Seung Shin Park, Jung Hee Kim
Endocrinology and Metabolism.2023; 38(4): 373. CrossRef - LC-MS based simultaneous profiling of adrenal hormones of steroids, catecholamines, and metanephrines
Jongsung Noh, Chaelin Lee, Jung Hee Kim, Seung Woon Myung, Man Ho Choi
Journal of Lipid Research.2023; 64(11): 100453. CrossRef - 2023 Korean Endocrine Society Consensus Guidelines for the Diagnosis and Management of Primary Aldosteronism
Jeonghoon Ha, Jung Hwan Park, Kyoung Jin Kim, Jung Hee Kim, Kyong Yeun Jung, Jeongmin Lee, Jong Han Choi, Seung Hun Lee, Namki Hong, Jung Soo Lim, Byung Kwan Park, Jung-Han Kim, Kyeong Cheon Jung, Jooyoung Cho, Mi-kyung Kim, Choon Hee Chung
Endocrinology and Metabolism.2023; 38(6): 597. CrossRef - Toward Systems-Level Metabolic Analysis in Endocrine Disorders and Cancer
Aliya Lakhani, Da Hyun Kang, Yea Eun Kang, Junyoung O. Park
Endocrinology and Metabolism.2023; 38(6): 619. CrossRef - Prevalence and Characteristics of Adrenal Tumors in an Unselected Screening Population
Ying Jing, Jinbo Hu, Rong Luo, Yun Mao, Zhixiao Luo, Mingjun Zhang, Jun Yang, Ying Song, Zhengping Feng, Zhihong Wang, Qingfeng Cheng, Linqiang Ma, Yi Yang, Li Zhong, Zhipeng Du, Yue Wang, Ting Luo, Wenwen He, Yue Sun, Fajin Lv, Qifu Li, Shumin Yang
Annals of Internal Medicine.2022; 175(10): 1383. CrossRef

 KES
KES
 PubReader
PubReader ePub Link
ePub Link Cite
Cite