Articles
- Page Path
- HOME > Endocrinol Metab > Volume 33(1); 2018 > Article
-
Original ArticleComparison of Immunohistochemistry and Direct Sanger Sequencing for Detection of the BRAFV600E Mutation in Thyroid Neoplasm
 Audioslide
Audioslide -
Hye-Seon Oh1, Hyemi Kwon1,2, Suyeon Park1, Mijin Kim1, Min Ji Jeon1, Tae Yong Kim1, Young Kee Shong1, Won Bae Kim1, Jene Choi3, Won Gu Kim1
 , Dong Eun Song3
, Dong Eun Song3
-
Endocrinology and Metabolism 2018;33(1):62-69.
DOI: https://doi.org/10.3803/EnM.2018.33.1.62
Published online: January 30, 2018
1Division of Endocrinology and Metabolism, Department of Internal Medicine, Asan Medical Center, University of Ulsan College of Medicine, Seoul, Korea.
2Division of Endocrinology and Metabolism, Kangbuk Samsung Hospital, Sungkyunkwan University School of Medicine, Seoul, Korea.
3Department of Pathology, Asan Medical Center, University of Ulsan College of Medicine, Seoul, Korea.
- Corresponding authors: Won Gu Kim. Division of Endocrinology and Metabolism, Department of Internal Medicine, Asan Medical Center, University of Ulsan College of Medicine, 88 Olympic-ro 43-gil, Songpa-gu, Seoul 05505, Korea. Tel: +82-2-3010-5883, Fax: +82-2-3010-6962, wongukim@amc.seoul.kr
- Corresponding authors: Dong Eun Song. Department of Pathology, Asan Medical Center, University of Ulsan College of Medicine, 88 Olympic-ro 43-gil, Songpa-gu, Seoul 05505, Korea. Tel: +82-2-3010-5998, Fax: +82-2-3010-7898, hipuha@hanmail.net
Copyright © 2018 Korean Endocrine Society
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
ABSTRACT
-
Background
- The BRAFV600E mutation is the most common genetic alteration identified in papillary thyroid carcinoma (PTC). Because of its costs effectiveness and sensitivity, direct Sanger sequencing has several limitations. The aim of this study was to evaluate the efficiency of immunohistochemistry (IHC) as an alternative method to detect the BRAFV600E mutation in preoperative and postoperative tissue samples.
-
Methods
- We evaluated 71 patients who underwent thyroid surgery with the result of direct sequencing of the BRAFV600E mutation. IHC staining of the BRAFV600E mutation was performed in 49 preoperative and 23 postoperative thyroid specimens.
-
Results
- Sixty-two patients (87.3%) had PTC, and of these, BRAFV600E was confirmed by direct sequencing in 57 patients (91.9%). In 23 postoperative tissue samples, the BRAFV600E mutation was detected in 16 samples (70%) by direct sequencing and 18 samples (78%) by IHC. In 24 fine needle aspiration (FNA) samples, BRAFV600E was detected in 18 samples (75%) by direct sequencing and 16 samples (67%) by IHC. In 25 core needle biopsy (CNB) samples, the BRAFV600E mutation was detected in 15 samples (60%) by direct sequencing and 16 samples (64%) by IHC. The sensitivity and specificity of IHC for detecting the BRAFV600E mutation were 77.8% and 66.7% in FNA samples and 99.3% and 80.0% in CNB samples.
-
Conclusion
- IHC could be an alternative method to direct Sanger sequencing for BRAFV600E mutation detection both in postoperative and preoperative samples. However, application of IHC to detect the BRAFV600E mutation in FNA samples is of limited value compared with direct sequencing.
- The BRAFV600E mutation is the most common mutation detected in papillary thyroid carcinoma (PTC). In Korea, PTCs harboring BRAFV600E were reported in up to 80% of the population [123]. The BRAF oncogene encodes the human gene for B-type Raf kinase. BRAFV600E is a T1799A point mutation in exon 15, resulting in the substitution of valine to glutamate at residue 600 (V600E). It activates the RAS/RAF/MAPK signaling pathway by disrupting hydrophobic interactions between residues in both the activation loop and the ATP-binding site [4]. BRAFV600E blocks apoptosis and regulates proliferation and invasiveness [5]. In addition, BRAFV600E is associated with aggressive clinical behavior and a higher risk of recurrence of PTC [678] and may help in detecting malignancy in some thyroid nodules [910]. The BRAFV600E mutation is one of the most stressed molecular markers for its clinical role in thyroid cancer.
- The gold standard method for molecular diagnosis of the BRAFV600E mutation is direct Sanger sequencing, which is highly reliable and specific [11]. However, the sensitivity of Sanger sequencing was reported to identify only 7% to 20% of mutated alleles in a background of wild-type alleles [1112]. The tumor content of malignant cells is reported to influence the sensitivity of Sanger sequencing [13]. Over the last decade, several alternatives to Sanger sequencing for detecting the BRAFV600E mutation have been reported with different sensitivities and specificities [11141516].
- Recently, immunohistochemistry (IHC) using a BRAFV600E-specific antibody (VE1) has been used to detect the BRAFV600E mutation [17]. IHC is a widely available and low-cost technique. Previous studies suggest that IHC is highly sensitive for detecting the BRAFV600E mutation [181920].
- In the present study, the efficiency of BRAFV600E IHC as an alternative method to direct Sanger sequencing was evaluated in preoperative and postoperative thyroid tissue samples.
INTRODUCTION
- Tissue samples and study design
- Seventy-one patients who underwent thyroid surgery between January 2011 and January 2013 in the Asan Medical Center, Seoul, Korea, were included in this retrospective study. Results on the BRAF mutation status using direct Sanger sequencing were available. Diagnostic preoperative tissue samples were obtained by either fine needle aspiration (FNA) or core needle biopsy (CNB). Postoperative formalin-fixed, paraffin-embedded (FFPE) samples were obtained after thyroid surgery. IHC staining for the BRAFV600E mutation was performed in preoperative and postoperative thyroid specimens. The results of BRAFV600E IHC were compared with those of direct Sanger sequencing. The study protocol was approved by the Institutional Review Board of the Asan Medical Center (AMC 2016-0821). Informed consent was obtained from all patients for BRAFV600E mutation analysis.
- Cytological and histopathological diagnosis
- All thyroid specimens were reviewed by an experienced endocrine pathologist (D.E.S.). FNA cytology results were categorized into six categories using the Bethesda system [21]. Preoperative CNB specimens were evaluated based on the criteria proposed by the Korean endocrine pathology thyroid CNB study group [22]. Surgical specimens were reviewed and diagnosed based on the World Health Organization classification criteria [23].
- Ultrasonography-guided fine needle aspiration and core needle biopsy procedures
- Neck ultrasound (US) examinations were performed using either an iU22 unit (Philips Healthcare, Bothell, WA, USA) or an EUB-7500 unit (Hitachi Medical Systems, Tokyo, Japan) equipped with a linear high-frequency probe (5 to 14 MHz), as previously reported [24]. The scanning protocol included both transverse and longitudinal real-time imaging. All US examinations and FNA/CNB procedures were performed by experienced radiologists. FNA procedures were performed under US guidance using a 23-gauge needle connected to a 10 mL syringe [25]. US-guided CNB procedures were performed using an 18-gauge, 1.1 or 1.6 cm excursion, double-action, spring-activated needle (TSK Ace-cut, Create Medic, Yokohama, Japan) [2627].
- Immunohistochemistry for the BRAFV600E mutation
- IHC for the BRAFV600E mutation was performed from 49 preoperative FNA cell blocks or CNB specimens and all postoperative thyroidectomy specimens using mouse anti-BRAF V600E (clone VE1, 1:4 dilution, Ventana Medical Systems, Tucson, AZ, USA) and the OptiView DAB IHC Detection Kit (Roche, Tucson, AZ, USA) on a Ventana BenchMark XT autoimmunostainer (Ventana Medical Systems) [28]. Whole tissue sections (4 µm) were transferred to poly-L-lysine-coated adhesive slides and dried at room temperature for 10 minutes. Antigen retrieval was performed using the heat-induced epitope retrieval method, followed by 20 minutes at 65℃ in an incubator. After incubation in CC1 solution (Roche) for 32 minutes, primary antibody was added and incubated for 16 minutes. A haptenated secondary antibody was added and incubated for a further 10 minutes. Alpha-hapten-horseradish peroxidase was added to the sections. After incubation for 10 minutes, diaminobenzidine was added and incubated for 10 minutes. Negative controls were performed by omitting the primary antibody. Positive controls were melanoma tissue.
- Slides were evaluated by an experienced pathologist (D.E.S.) who was blinded to the BRAF mutational status. Diffuse homogeneous cytoplasmic staining in all tumor cells was considered as positive. Non-specific staining of colloids and equivocally weak or focal cytoplasmic staining was considered as negative.
- Direct Sanger sequencing for BRAF
- Sanger sequencing was performed as previously described [27]. Briefly, exon 15 of the BRAF gene was amplified by polymerase chain reaction (PCR) using the primers 5′-TGCTTGCTCTGATAGGAAAATG-3′ (forward) and 5′-CTGATGGG ACCCACTCC AT-3′ (reverse). The amplification protocol comprised initial denaturation at 94℃ for 2 minutes; 40 cycles of denaturation at 95℃ for 10 seconds, annealing at 60℃ for 30 seconds, and extension at 68℃ for 40 seconds; followed by a final extension at 68℃ for 7 minutes using KOD FX Taq DNA polymerase (Toyobo, Osaka, Japan). PCR products were electrophoresed on 2.5% (wt/vol) agarose gels and subsequently sequenced using the above forward primer and Big Dye Terminator (ABI Systems, Applied Biosystems, Foster City, CA, USA). DNA sequences and the BRAF mutation were determined using an ABI-PRISM 3100 automatic sequencer (Applied Biosystems).
- Statistical analysis
- Statistical analyses were performed using R version 3.0 and the R libraries (R Foundation for Statistical Computing, Vienna, Austria, http://www.R-project.org). Categorical variables are presented as numbers with percentages. McNemar's chi-square test was used to evaluate the performance of two tests. All P values were two-sided and P values <0.05 were considered statistically significant.
METHODS
- Baseline characteristics of study subjects and tissue samples
- Sixty-two of 71 patients (87.3%) had PTC including one patient with a follicular variant of PTC, and of these, the BRAFV600E mutation was detected in 57 patients (91.9%) by direct sequencing. Four patients (5.6%) had follicular thyroid carcinoma, and an additional four patients (5.6%) had nodular hyperplasia. Wild-type BRAF was confirmed in 14 tumors (19.7%).
- Comparison of direct Sanger sequencing and IHC results for the BRAFV600E mutation using postoperative samples
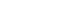
- Table 1 shows the results of direct Sanger sequencing and IHC staining for the BRAFV600E mutation in 23 postoperative FFPE samples. The BRAFV600E mutation was detected in 16 samples (70%) using direct sequencing. IHC for BRAFV600E was positive in 18 FFPE samples (78%). Of these samples, two showed discrepant results between the two methods. IHC results were positive, whereas the results of direct sequencing showed wild-type BRAF (Fig. 1A, B). The sensitivity and specificity of IHC for detecting the BRAFV600E mutation in postoperative samples were 100% and 71.4%. There was no significant difference in detecting the BRAFV600E mutation between two tests in postoperative samples (P=0.480).
- Comparison of direct Sanger sequencing and IHC results for the BRAFV600E mutation using preoperative samples
- Table 2 shows the results of direct Sanger sequencing and IHC staining for the BRAFV600E mutation in 49 preoperative samples including 24 FNA samples and 25 CNB samples. The BRAFV600E mutation was detected in 33 samples (67%) using direct sequencing and 32 samples (65%) using IHC. In preoperative samples, inconsistent results between the two methods were detected in nine patients (18.4%). Of these, four were positive by IHC and negative by direct sequencing. Five patients were negative with IHC and positive with direct sequencing methods. The sensitivity and specificity of IHC for detecting the BRAFV600E mutation in preoperative samples were 84.9% and 75.0%. No difference was found between two tests in detecting the BRAFV600E mutation in preoperative samples (P=0.999).
- In 24 FNA specimens, the BRAFV600E mutation was detected in 18 samples (75%) by direct sequencing and 16 samples (67%) by IHC. The results of the two methods were discordant in six of 24 samples (25%). Two samples were positive for the BRAFV600E mutation by IHC, but direct sequencing results showed wild-type BRAF. The BRAFV600E mutation was detected in four samples using direct sequencing, but these were negative using IHC. BRAFV600E mutations from two samples using direct sequencing are shown in Fig. 1C, D. However, results of IHC were negative in the two corresponding FNA cell blocks as shown in Fig. 1E, F. The sensitivity and specificity of IHC for detecting the BRAFV600E mutation in FNA samples were 77.8% and 66.7%. There was no statistical significant difference in detecting the BRAFV600E mutation between two tests in FNA samples (P=0.683).
- In 25 CNB specimens, the BRAFV600E mutation was detected in 15 samples (60%) by direct sequencing and 16 samples (64%) by IHC. The results of direct sequencing and IHC differed in three of 25 samples (12%). The BRAFV600E mutation was detected in one sample by direct sequencing, but was negative in IHC. Two samples were positive for the BRAFV600E mutation using IHC, but were not detected by direct sequencing (Fig. 1G, H). The sensitivity and specificity of IHC for detecting the BRAFV600E mutation in CNB samples were 99.3% and 80.0%. There was no significant difference in detecting the BRAFV600E mutation between two tests in CNB samples (P=0.999).
RESULTS
- The presence of the BRAFV600E mutation is considered an intermediate risk factor in the recently revised American Thyroid Association guideline for thyroid nodules and differentiated thyroid cancer [29]. This indicates that the presence of BRAFV600E is generally considered as a risk factor for poor prognosis or aggressive behavior of thyroid cancer. BRAFV600E mutation analysis is difficult to perform in every thyroid cancer patient, because of the issue of cost-effectiveness.
- The efficiency of IHC, a low-cost method, for detecting the BRAFV600E mutation was evaluated as an alternative technique to direct Sanger sequencing in different preoperative and postoperative thyroid tissue samples. The sensitivity of IHC staining for detecting the BRAFV600E mutation was found to be superior to that of direct Sanger sequencing in postoperative tissue samples. In preoperative tissue samples, IHC staining performed better in CNB samples.
- The BRAFV600E mutation was found in 92% of PTC patients in the present study. This result is consistent with findings of previous studies demonstrating a relatively high prevalence of the BRAFV600E mutation in Korean PTC patients [130]. This implies that detection of the BRAFV600E mutation could play a role in the diagnosis or management of PTC patients, especially in Korea.
- According to our data, the concordance between direct Sanger sequencing and IHC for detecting BRAFV600E was superior in postoperative tissue samples compared with preoperative tissue samples. Discordance between the two methods in postoperative tissue samples was found in only two patients, who were wild-type by direct Sanger sequencing and positive using IHC. In preoperative tissue samples, discordant results between the two methods were found in nine samples, of which four were wild-type by direct Sanger sequencing and positive using IHC, and the remaining five were vice versa. These results imply that IHC is more suitable for postoperative tissue samples than preoperative tissue samples.
- Preoperative tissue samples were obtained by FNA and CNB. Data showed that CNB samples were more suitable for IHC staining than FNA samples, even though there were no statistical differences between Sanger sequencing and IHC staining methods on both tissue samples. According to previous studies, IHC on FNA samples shows relatively less specificity and sensitivity than on FFPE samples [183132]. However, Crescenzi et al. [33] reported that IHC performed on thyroid CNB samples perfectly matched genetic analysis of BRAF status. These data support the use of IHC on CNB samples to detect BRAFV600E at preoperative evaluation of thyroid nodules.
- Five tissue samples were positive by direct Sanger sequencing and negative by IHC, with four FNA samples and one CNB sample. All of these samples were collected preoperatively. These findings may be due to loss of antigen expression of the mutation. For example, tissue ischemic areas surrounding necrotic areas or damaged tissue have reduced BRAFV600E protein expression, which may be the cause of the false-negative IHC results, particularly in small biopsies such as FNA [1834].
- As shown in Supplemental Table S1, the FNA sample from subject number 1 showed negative results by direct Sanger sequencing and positive results by IHC. In this patient, subsequent reverse transcriptase-PCR revealed the BRAFV600E mutation. This implies that methods for detecting the BRAFV600E mutation may affect the results of mutation analysis. Combination of IHC and molecular approaches has been suggested for better detection of the BRAFV600E mutation [35]. However, the cost-effectiveness of using these approaches requires consideration.
- In conclusion, IHC could be an alternative approach to direct Sanger sequencing for detecting the BRAFV600E mutation both in postoperative and preoperative tissue samples. However, application of IHC in FNA samples was of limited value compared with direct sequencing. Direct Sanger sequencing is the gold standard method for detecting the BRAFV600E mutation and is highly specific. Disadvantages include its relative low sensitivity compared with other tests [1135] and high costs. IHC on appropriate tissue samples is an alternative and superior approach in detecting the BRAFV600E mutation because of its improved performance and low cost.
DISCUSSION
-
Acknowledgements
- This study was supported by the National Research Foundation (NRF) of Korea Research Grant (NRF-2015R1C1A1A02036597).
ACKNOWLEDGMENTS
-
CONFLICTS OF INTEREST: No potential conflict of interest relevant to this article was reported.
-
AUTHOR CONTRIBUTIONS: Conception or design: Y.K.S., W.B.K., W.G.K., D.E.S. Acquisition of data: H.S.O., H.K., P.S., M.K., J.C. Analysis or interpretation of data: H.S.O., H.K., M.J.J., T.Y.K., J.C., W.G.K. Drafting the work or revising: H.S.O., H.K., W.G.K., D.E.S. Final approval of the manuscript: Y.K.S., W.B.K., W.G.K., D.E.S.
Article information
SUPPLEMENTARY MATERIAL
Supplemental Table S1
- 1. Kim KH, Kang DW, Kim SH, Seong IO, Kang DY. Mutations of the BRAF gene in papillary thyroid carcinoma in a Korean population. Yonsei Med J 2004;45:818–821. ArticlePubMed
- 2. Kim TY, Kim WB, Song JY, Rhee YS, Gong G, Cho YM, et al. The BRAF mutation is not associated with poor prognostic factors in Korean patients with conventional papillary thyroid microcarcinoma. Clin Endocrinol (Oxf) 2005;63:588–593. ArticlePubMed
- 3. Jung CK, Im SY, Kang YJ, Lee H, Jung ES, Kang CS, et al. Mutational patterns and novel mutations of the BRAF gene in a large cohort of Korean patients with papillary thyroid carcinoma. Thyroid 2012;22:791–797. ArticlePubMed
- 4. Wan PT, Garnett MJ, Roe SM, Lee S, Niculescu-Duvaz D, Good VM, et al. Mechanism of activation of the RAF-ERK signaling pathway by oncogenic mutations of B-RAF. Cell 2004;116:855–867. ArticlePubMed
- 5. Nikiforov YE, Nikiforova MN. Molecular genetics and diagnosis of thyroid cancer. Nat Rev Endocrinol 2011;7:569–580. ArticlePubMedPDF
- 6. Tufano RP, Teixeira GV, Bishop J, Carson KA, Xing M. BRAF mutation in papillary thyroid cancer and its value in tailoring initial treatment: a systematic review and meta-analysis. Medicine (Baltimore) 2012;91:274–286. ArticlePubMed
- 7. Xing M, Westra WH, Tufano RP, Cohen Y, Rosenbaum E, Rhoden KJ, et al. BRAF mutation predicts a poorer clinical prognosis for papillary thyroid cancer. J Clin Endocrinol Metab 2005;90:6373–6379. ArticlePubMed
- 8. Elisei R, Viola D, Torregrossa L, Giannini R, Romei C, Ugolini C, et al. The BRAF(V600E) mutation is an independent, poor prognostic factor for the outcome of patients with low-risk intrathyroid papillary thyroid carcinoma: single-institution results from a large cohort study. J Clin Endocrinol Metab 2012;97:4390–4398. ArticlePubMedPDF
- 9. Choi SH, Baek JH, Lee JH, Choi YJ, Song DE, Chung KW, et al. Evaluation of the clinical usefulness of BRAFV600E mutation analysis of core-needle biopsy specimens in thyroid nodules with previous atypia of undetermined significance or follicular lesions of undetermined significance results. Thyroid 2015;25:897–903. ArticlePubMed
- 10. Borrelli N, Ugolini C, Giannini R, Antonelli A, Giordano M, Sensi E, et al. Role of gene expression profiling in defining indeterminate thyroid nodules in addition to BRAF analysis. Cancer Cytopathol 2016;124:340–349. ArticlePubMed
- 11. Ihle MA, Fassunke J, Konig K, Grunewald I, Schlaak M, Kreuzberg N, et al. Comparison of high resolution melting analysis, pyrosequencing, next generation sequencing and immunohistochemistry to conventional Sanger sequencing for the detection of p.V600E and non-p.V600E BRAF mutations. BMC Cancer 2014;14:13ArticlePubMedPMCPDF
- 12. Monzon FA, Ogino S, Hammond ME, Halling KC, Bloom KJ, Nikiforova MN. The role of KRAS mutation testing in the management of patients with metastatic colorectal cancer. Arch Pathol Lab Med 2009;133:1600–1606. ArticlePubMedPDF
- 13. Tol J, Dijkstra JR, Vink-Borger ME, Nagtegaal ID, Punt CJ, Van Krieken JH, et al. High sensitivity of both sequencing and real-time PCR analysis of KRAS mutations in colorectal cancer tissue. J Cell Mol Med 2010;14:2122–2131. ArticlePubMed
- 14. Colomba E, Helias-Rodzewicz Z, Von Deimling A, Marin C, Terrones N, Pechaud D, et al. Detection of BRAF p.V600E mutations in melanomas: comparison of four methods argues for sequential use of immunohistochemistry and pyrosequencing. J Mol Diagn 2013;15:94–100. ArticlePubMed
- 15. Carbonell P, Turpin MC, Torres-Moreno D, Molina-Martinez I, Garcia-Solano J, Perez-Guillermo M, et al. Comparison of allelic discrimination by dHPLC, HRM, and TaqMan in the detection of BRAF mutation V600E. J Mol Diagn 2011;13:467–473. ArticlePubMedPMC
- 16. Jiang W, Wang W, Fu F, Teng X, Wang H, Wang H, et al. A more sensitive platform for the detection of low-abundance BRAF(V600E) mutations. Mol Cell Biochem 2012;366:49–58. ArticlePubMedPDF
- 17. Routhier CA, Mochel MC, Lynch K, Dias-Santagata D, Louis DN, Hoang MP. Comparison of 2 monoclonal antibodies for immunohistochemical detection of BRAF V600E mutation in malignant melanoma, pulmonary carcinoma, gastrointestinal carcinoma, thyroid carcinoma, and gliomas. Hum Pathol 2013;44:2563–2570. ArticlePubMed
- 18. Dvorak K, Aggeler B, Palting J, McKelvie P, Ruszkiewicz A, Waring P. Immunohistochemistry with the anti-BRAF V600E (VE1) antibody: impact of pre-analytical conditions and concordance with DNA sequencing in colorectal and papillary thyroid carcinoma. Pathology 2014;46:509–517. ArticlePubMedPMC
- 19. Bullock M, O'Neill C, Chou A, Clarkson A, Dodds T, Toon C, et al. Utilization of a MAB for BRAF(V600E) detection in papillary thyroid carcinoma. Endocr Relat Cancer 2012;19:779–784. ArticlePubMed
- 20. Rossi ED, Martini M, Capodimonti S, Cenci T, Straccia P, Angrisani B, et al. Analysis of immunocytochemical and molecular BRAF expression in thyroid carcinomas: a cytohistologic institutional experience. Cancer Cytopathol 2014;122:527–535. ArticlePubMed
- 21. Cibas ES, Ali SZ. The Bethesda System for reporting thyroid cytopathology. Thyroid 2009;19:1159–1165. ArticlePubMed
- 22. Jung CK, Min HS, Park HJ, Song DE, Kim JH, Park SY, et al. Pathology reporting of thyroid core needle biopsy: a proposal of the Korean endocrine pathology thyroid core needle biopsy study group. J Pathol Transl Med 2015;49:288–299. ArticlePubMedPMCPDF
- 23. DeLellis RA. Pathology and genetics of tumours of endocrine organs; Lyon: IARC Press; 2004.
- 24. Oh HS, Kim WG, Park S, Kim M, Kwon H, Jeon MJ, et al. Serial neck ultrasonographic evaluation of changes in papillary thyroid carcinoma during pregnancy. Thyroid 2017;27:773–777. ArticlePubMed
- 25. Kwon H, Kim WG, Eszlinger M, Paschke R, Song DE, Kim M, et al. Molecular diagnosis using residual liquid-based cytology materials for patients with nondiagnostic or indeterminate thyroid nodules. Endocrinol Metab (Seoul) 2016;31:586–591. ArticlePubMedPMC
- 26. Jeon MJ, Song DE, Jung CK, Kim WG, Kwon H, Lee YM, et al. Impact of reclassification on thyroid nodules with architectural atypia: from non-invasive encapsulated follicular variant papillary thyroid carcinomas to non-invasive follicular thyroid neoplasm with papillary-like nuclear features. PLoS One 2016;11:e0167756ArticlePubMedPMC
- 27. Choi SH, Baek JH, Lee JH, Choi YJ, Ha EJ, Song DE, et al. Initial clinical experience with BRAF(V600E) mutation analysis of core-needle biopsy specimens from thyroid nodules. Clin Endocrinol (Oxf) 2016;84:607–613. ArticlePubMed
- 28. Koperek O, Kornauth C, Capper D, Berghoff AS, Asari R, Niederle B, et al. Immunohistochemical detection of the BRAF V600E-mutated protein in papillary thyroid carcinoma. Am J Surg Pathol 2012;36:844–850. ArticlePubMed
- 29. Haugen BR, Alexander EK, Bible KC, Doherty GM, Mandel SJ, Nikiforov YE, et al. 2015 American Thyroid Association management guidelines for adult patients with thyroid nodules and differentiated thyroid cancer: the American Thyroid Association Guidelines Task Force on Thyroid Nodules and Differentiated Thyroid Cancer. Thyroid 2016;26:1–133. ArticlePubMedPMC
- 30. Lee SE, Hwang TS, Choi YL, Kim WY, Han HS, Lim SD, et al. Molecular profiling of papillary thyroid carcinoma in Korea with a high prevalence of BRAF(V600E) mutation. Thyroid 2017;27:802–810. ArticlePubMed
- 31. Leslie C, Grieu-Iacopetta F, Richter A, Platten M, Murray J, Frost FA, et al. BRAF p.Val600Glu (V600E) mutation detection in thyroid fine needle aspiration cell block samples: a feasibility study. Pathology 2015;47:432–438. ArticlePubMed
- 32. Wobker SE, Kim LT, Hackman TG, Dodd LG. Use of BRAF v600e immunocytochemistry on FNA direct smears of papillary thyroid carcinoma. Cancer Cytopathol 2015;123:531–539. ArticlePubMed
- 33. Crescenzi A, Guidobaldi L, Nasrollah N, Taccogna S, Cicciarella Modica DD, Turrini L, et al. Immunohistochemistry for BRAF(V600E) antibody VE1 performed in core needle biopsy samples identifies mutated papillary thyroid cancers. Horm Metab Res 2014;46:370–374. ArticlePubMedPDF
- 34. Capper D, Preusser M, Habel A, Sahm F, Ackermann U, Schindler G, et al. Assessment of BRAF V600E mutation status by immunohistochemistry with a mutation-specific monoclonal antibody. Acta Neuropathol 2011;122:11–19. ArticlePubMedPDF
- 35. Martinuzzi C, Pastorino L, Andreotti V, Garuti A, Minuto M, Fiocca R, et al. A combination of immunohistochemistry and molecular approaches improves highly sensitive detection of BRAF mutations in papillary thyroid cancer. Endocrine 2016;53:672–680. ArticlePubMedPDF
References
Detection of the BRAFV600E mutation in different thyroid tissue specimens with inconsistent results between direct Sanger sequencing and immunohistochemistry (IHC) (A–F). Two postoperative thyroidectomy specimens were positive by immunohistochemistry for the BRAFV600E mutation (A, B, ×400), but negative by direct Sanger sequencing. Two preoperative fine needle aspiration cell block specimens showed discrepant results with positive direct Sanger sequencing (C, D) results and negative IHC results (E, F, ×400). Two preoperative core needle biopsy specimens were positive by IHC for the BRAFV600E mutation (G, H, ×400), but negative by direct Sanger sequencing.
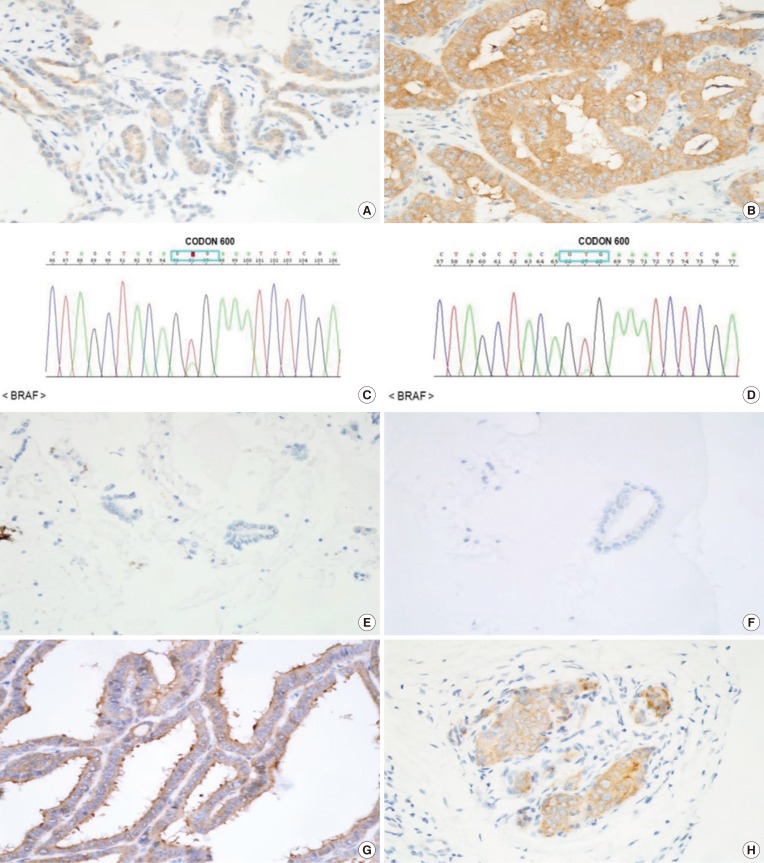
Comparison of the Results of Direct Sanger Sequencing and IHC for BRAFV600E in Postoperative Samples
| BRAFV600E IHC | Sanger sequencing | Total | |
|---|---|---|---|
| Wild-type | BRAFV600E | ||
| Negative | 5 (71) | 0 | 5 (22) |
| Positive | 2 (29) | 16 (100) | 18 (78) |
| Total | 7 (30) | 16 (70) | 23 (100) |
Comparison of the Results of Direct Sanger Sequencing and IHC for BRAFV600E in Preoperative Samples
Figure & Data
References
Citations

- Circulating Nucleic Acids in Colorectal Cancer: Diagnostic and Prognostic Value
Somayeh Igder, Mozhdeh Zamani, Shima Fakher, Morvarid Siri, Hassan Ashktorab, Negar Azarpira, Pooneh Mokarram, Sowjanya Thatikonda
Disease Markers.2024; 2024: 1. CrossRef - The Accurate Interpretation and Clinical Significance of Morphological Features of Fine Needle Aspiration Cells in Papillary Thyroid Carcinoma
Xue-Jiao Xiong, Ming-Ming Xiao, Yi-Xia Zhang, Dong-Ge Liu, Mu-Lan Jin, Jian Wang, Hong-Tao Xu, Qing-Chang Li, Guang-Ping Wu, Giovanni Tuccari
Analytical Cellular Pathology.2023; 2023: 1. CrossRef - An effective approach for BRAF V600E mutation analysis of routine thyroid fine needle aspirates
Tanupriya Agrawal, Liqiang Xi, Winnifred Navarro, Mark Raffeld, Snehal B. Patel, Mark J. Roth, Joanna Klubo‐Gwiezdzinska, Armando C. Filie
Cytopathology.2022; 33(3): 344. CrossRef - A dual identification strategy based on padlock ligation and CRISPR/Cas14a for highly specific detection of BRAF V600E mutation in clinical samples
Weicheng Shi, Yao Gong, Decai Zhang, Tiantian Yang, Ming Yi, Jingyi Tan, Shijia Ding, Wei Cheng
Analytical Methods.2022; 14(19): 1913. CrossRef - Research Progress of BRAF V600E Gene Mutation in Papillary Thyroid Carcinoma
延泽 刘
Advances in Clinical Medicine.2022; 12(09): 8499. CrossRef - VE1 immunohistochemistry is an adjunct tool for detection of BRAFV600E mutation: Validation in thyroid cancer patients
Faiza A. Rashid, Sobia Tabassum, Mosin S. Khan, Hifzur R. Ansari, Muhammad Asif, Ahmareen K. Sheikh, Syed Sameer Aga
Journal of Clinical Laboratory Analysis.2021;[Epub] CrossRef - BRAF testing in a South African cohort of MLH1 deficient endometrial carcinomas: lessons learnt
Reubina Wadee, Wayne Grayson
Southern African Journal of Gynaecological Oncology.2021; 13(1): 1. CrossRef - Association between mutation profiles and clinicopathological features in Chinese patients with thyroid cancer
Changwen Jing, Haixia Cao, Rong Ma, Jianzhong Wu, Zhuo Wang
Precision Medical Sciences.2021; 10(3): 113. CrossRef - Development of a Molecular Assay for Detection and Quantification of theBRAFVariation in Residual Tissue From Thyroid Nodule Fine-Needle Aspiration Biopsy Specimens
Guodong Fu, Ronald S. Chazen, Christina MacMillan, Ian J. Witterick
JAMA Network Open.2021; 4(10): e2127243. CrossRef - Variations in MAP kinase gladiators and risk of differentiated thyroid carcinoma
Faiza Rashid, Ghulam Bhat, Mosin Khan, Sobia Tabassum, Mohammad Bhat
Molecular and Clinical Oncology.2021;[Epub] CrossRef - Порівняльне імуногістохімічне дослідження BRAFV600E-позитивних і BRAFV600E-негативних радіогенних і спорадичних папілярних тиреоїдних карцином
L. Yu. Zurnadzhy, T.I. Rogounovitch, V.O. Saenko, M.Yu. Bolgov, S.V. Masiuk, S.V. Burko, T.L. Degtyaryova, S.V. Chernyshov, S.V. Gulevatyi, N. Mitsutake, M.D. Tronko, T.I. Bogdanova
Endokrynologia.2021; 26(2): 105. CrossRef - Evaluation of the expression levels of BRAFV600E mRNA in primary tumors of thyroid cancer using an ultrasensitive mutation assay
Tien Viet Tran, Kien Xuan Dang, Quynh Huong Pham, Ung Dinh Nguyen, Nhung Thi Trang Trinh, Luong Van Hoang, Son Anh Ho, Ba Van Nguyen, Duc Trong Nguyen, Dung Tuan Trinh, Dung Ngoc Tran, Arto Orpana, Ulf-Håkan Stenman, Jakob Stenman, Tho Huu Ho
BMC Cancer.2020;[Epub] CrossRef - VE1 Immunohistochemistry Improves the Limit of Genotyping for Detecting BRAFV600E Mutation in Papillary Thyroid Cancer
Sonam Choden, Somboon Keelawat, Chan Kwon Jung, Andrey Bychkov
Cancers.2020; 12(3): 596. CrossRef - Comparison of Molecular Methods and BRAF Immunohistochemistry (VE1 Clone) for the Detection of BRAF V600E Mutation in Papillary Thyroid Carcinoma: A Meta-Analysis
Kyle G. Parker, Michael G. White, Nicole A. Cipriani
Head and Neck Pathology.2020; 14(4): 1067. CrossRef - Next generation sequencing based detection of 15 target genes mutations in papillary thyroid carcinoma
Zhuo Wang, Changwen Jing, Haixia Cao, SiWen Liu, Jianzhong Wu, Rong Ma
Precision Medical Sciences.2020; 9(2): 90. CrossRef - A Systematic Review and Meta-Analysis of the Diagnostic Performance of BRAF V600E Immunohistochemistry in Thyroid Histopathology
Ranjit Singarayer, Ozgur Mete, Laure Perrier, Lehana Thabane, Sylvia L. Asa, Stan Van Uum, Shereen Ezzat, David P. Goldstein, Anna M. Sawka
Endocrine Pathology.2019; 30(3): 201. CrossRef - Comparison of droplet digital PCR and direct Sanger sequencing for the detection of the BRAFV600E mutation in papillary thyroid carcinoma
Zhuo Wang, Kejing Sun, Changwen Jing, Haixia Cao, Rong Ma, Jianzhong Wu
Journal of Clinical Laboratory Analysis.2019;[Epub] CrossRef - Immunohistochemistry Innovations for Diagnosis and Tissue-Based Biomarker Detection
Narittee Sukswai, Joseph D. Khoury
Current Hematologic Malignancy Reports.2019; 14(5): 368. CrossRef - Immunohistochemistry is a feasible method to screen BRAF V600E mutation in colorectal and papillary thyroid carcinoma
Xiangyan Zhang, Lili Wang, Jigang Wang, Han Zhao, Jie Wu, Shuhong Liu, Lu Zhang, Yujun Li, Xiaoming Xing
Experimental and Molecular Pathology.2018; 105(1): 153. CrossRef

 KES
KES
 PubReader
PubReader ePub Link
ePub Link Cite
Cite