Roles of Protein Arginine Methyltransferases in the Control of Glucose Metabolism
Article information
Abstract
Glucose homeostasis is tightly controlled by the regulation of glucose production in the liver and glucose uptake into peripheral tissues, such as skeletal muscle and adipose tissue. Under prolonged fasting, hepatic gluconeogenesis is mainly responsible for glucose production in the liver, which is essential for tissues, organs, and cells, such as skeletal muscle, the brain, and red blood cells. Hepatic gluconeogenesis is controlled in part by the concerted actions of transcriptional regulators. Fasting signals are relayed by various intracellular enzymes, such as kinases, phosphatases, acetyltransferases, and deacetylases, which affect the transcriptional activity of transcription factors and transcriptional coactivators for gluconeogenic genes. Protein arginine methyltransferases (PRMTs) were recently added to the list of enzymes that are critical for regulating transcription in hepatic gluconeogenesis. In this review, we briefly discuss general aspects of PRMTs in the control of transcription. More specifically, we summarize the roles of four PRMTs: PRMT1, PRMT 4, PRMT 5, and PRMT 6, in the control of hepatic gluconeogenesis through specific regulation of FoxO1- and CREB-dependent transcriptional events.
INTRODUCTION
Protein arginine methyltransferases (PRMTs) comprise a group of enzymes that catalyze the transfer of methyl groups to the amino acid arginine or the arginine residues of proteins in eukaryotic species. In mammals, nine isoforms have been identified to date that can be subdivided into three major types: I, II, and III. While all three types of PRMTs are capable of catalyzing monomethylation of arginine residues, type I PRMTs can transfer a second methyl group to the monomethylated nitrogen atom of the guanidino group of monomethyl arginine, resulting in the formation of proteins containing one or more asymmetric dimethylarginine residues. Type II PRMTs transfer a methyl group to the unmodified nitrogen atom of the guanidine group of monomethyl arginine to form proteins with symmetric dimethylarginine residues. Type III PRMTs are a group of PRMTs that can only generate monomethylarginine [1,2].
Type I PRMTs comprise PRMT1, PRMT3, PRMT4, PRMT6, and PRMT8 and have been shown to function as transcriptional activators. In earlier reports, PRMTs promoted active transcription of target genes by mediating asymmetric dimethylation of arginine residues on histones, resulting in increased acetylation and the subsequent activation of transcription. Examples include arginine 3 methylation on histone H4 (H4R3) by PRMT1 and arginine 17 and 26 methylation on histone H3 (H3R17 and H3R26) by PRMT4 [3,4,5]. More recent reports suggest that PRMTs can affect transcription by direct modification of transcriptional regulators. Type II PRMTs (PRMT5, PRMT7, and PRMT9) are less well characterized and may function as transcriptional repressors, although some reports suggest that they can function as activators of transcription. PRMT5, a predominant type II PRMT in mammals, represses transcription by promoting symmetric dimethylation of arginine 8 on histone H3 (H3R8) and symmetric dimethylation of arginine 3 on histone H4 (H4R3) [6]. However, PRMT5 can also increase transcription under certain conditions [7,8]. The role of type III PRMTs in the control of transcription has not been well characterized to date.
Interestingly, recent reports suggest that PRMTs function as regulators of fuel metabolism through interaction with a number of transcriptional regulators in the pathway. In this review, we summarize recent findings regarding the roles of PRMT isoforms in the control of fuel metabolism, with emphasis on glucose metabolism. We restrict our discussions to four widely studied isoforms: PRMT1, PRMT4, PRMT5, and PRMT6.
PRMT1
PRMT1 is a type I PRMT that catalyzes the formation of asymmetric dimethylarginine. It is expressed ubiquitously in most tissues and may account for up to 85% of all PRMT activity in the cell depending on experimental conditions [9]. PRMT1 may regulate transcription by promoting asymmetric dimethylation of H4R3 [3]. In addition, it can also function as a coactivator of transcription factors by direct modification of arginine residues on the transcription factors themselves. For example, PRMT1-dependent methylation of RUNX1 is essential for its dissociation from mSin3A, resulting in transcription [10].
FoxO1 is a transcription factor containing a forkhead box domain that is conserved in various eukaryotic species. It is involved in the control of signaling pathways in cellular stress resistance, longevity, apoptosis, and metabolism. Normally, FoxO1 is present in the nucleus and functions as a transcriptional activator in the absence of growth factors or insulin. In the presence of such signals, three major serine/threonine residues (Thr24, Ser253, and Ser316 in Mus muculus) are phosphorylated by Akt or its related AGC kinases, resulting in an association with 14-3-3, nuclear exclusion, ubiquitination, and subsequent degradation by a proteasome-dependent pathway [11,12,13]. A recent study found that PRMT1 can also modulate FoxO1 activity by controlling its nuclear localization. In the presence of oxidative stress, PRMT1 levels are increased [14]. PRMT1 promotes asymmetric dimethylation of FoxO1 at arginines 248 and 250, thereby inhibiting phosphorylation of the adjacent serine 253, increasing its nuclear retention and promoting cell survival. Thus, PRMT1 may function as a transcriptional co-activator for FoxO1-dependent transcriptional events.
FoxO1 has been shown to be a major transcriptional activator for genes involved in hepatic gluconeogenesis. FoxO1 binding sites, termed insulin response elements, have been mapped to a number of genes in the pathway, including the genes encoding phosphoenolpyruvate carboxykinase (PEPCK), glucose 6-phosphatase (G6Pase), and peroxisome proliferator-activated receptor γ co-activator 1α (PGC-1α) [15,16]. Furthermore, transgenic expression of FoxO1 in the liver increased gluconeogenesis, while liver-specific depletion of FoxO1 in mice reduced glucose production in vivo, illustrating the critical role of this factor in hepatic gluconeogenesis [17,18].
Recent reports also confirmed that PRMT1 regulates FoxO1-dependent control of hepatic gluconeogenesis [19]. PRMT1 was shown to modulate asymmetric dimethylation of FoxO1 at arginines 248 and 250 in the mouse liver, disturbing the ability of insulin to reduce glucose production in the liver. Overexpression of PRMT1 promotes increased glucose production by hepatocytes, while acute depletion of PRMT1 in mice reduces hepatic gluconeogenesis by promoting increased phosphorylation of FoxO1 at serine 253, leading to the decreased expression of FoxO1 target genes during gluconeogenesis in mice. However, the precise role of PRMT1 in the control of FoxO1-dependent gluconeogenesis should be further delineated by using liver-specific PRMT1 knockout mice. In addition, further study is necessary to determine the mechanism underlying the control of PRMT1 activity in response to fasting or insulin resistance in the liver. We observed the unusual localization of PRMT1 in cytosolic stress granules upon serum starvation. Clarification regarding whether PRMT1 is also involved in the general control of posttranscriptional modification would be of interest.
PRMT4
PRMT4, also known as coactivator-associated arginine methyltransferase 1, is widely known as a coactivator for various nuclear hormone receptors, including androgen receptor, estrogen receptor, and thyroid hormone receptor [20,21]. It also exerts coactivator effects by interacting with other coactivators, including cAMP response element-binding protein (CREB) binding protein (CBP), P300/CBP-associated factor, steroid receptor coactivator-1 (SRC-1), and PRMT1, affecting their activity by direct methylation of their arginine residues in some cases or by modulating the arginine residues of histone H3 (H3R17 and H3R26) [4,5,22,23].
PRMT4 can function as an activator of myogenesis by acting as a coactivator of myocyte-specific enhancer factor 2C, nuclear receptor coactivator 3 [24]. As a coactivator for nuclear factor-κB, PRMT4 plays a role in the inflammatory stimulation of monocytes [25]. During adipogenesis, PRMT4 plays a role as a coactivator of PPAR-γ, promoting fat cell differentiation [26]. Interestingly, aberrant expression of PRMT4 was detected in prostate carcinoma, indicating the potential importance of this factor in normal signaling cascades in prostate cells [27]. In addition, PRMT4 may also be involved in the control of mRNA stability by methylation of HuR, an mRNA-stabilizing protein [28].
CREB plays a key role in the control of hepatic gluconeogenesis. Under fasting conditions, the pancreatic hormone glucagon activates cAMP signals in hepatocytes and activates protein kinase A (PKA). PKA promotes hepatic gluconeogenesis at the transcription level by promoting phosphorylation of CREB at serine 133. This in turn is crucial for recruiting coactivators such as CBP and p300, leading to the activation of target genes such as PEPCK, G6Pase, and PGC-1α. Indeed, liver-specific expression of A-CREB, a CREB inhibitor, or siRNA targeting CREB lowered blood glucose levels, a response associated with reduced gluconeogenic gene expression. This underscores the importance of this transcription factor in hepatic gluconeogenesis in vivo [29,30].
A recent report suggested that PRMT4 controls hepatic gluconeogenesis by interacting with CREB in hepatocytes. Krones-Herzig and colleague [31] showed that PRMT4 can associate physically with CREB on the promoters of its target genes, such as the PEPCK and G6Pase promoters. Recruited PRMT4 in turn prompts direct asymmetric dimethylation of histone H3 at arginine 2. This promotes increased histone H3 acetylation by CBP/p300, leading to the increased transcription of CREB target genes. This report, however, was somewhat contradictory to a previous report showing that PRMT4 turns off CREB-dependent transcription by selective arginine methylation in the KIX domain of CBP, a CREB-interacting domain, to dissociate it from CREB [32]. Further analysis is necessary to delineate the relative contributions of arginine methylation in CBP and histone H3 to CREB-dependent transcription.
PRMT5
PRMT5 is a type II PRMT that promotes symmetric dimethylation of arginine residues in substrates. Normally, symmetric dimethylation of arginine in histones is regarded as a marker for transcriptional inhibition. PRMT5 interacts with various transcriptional regulators, including Brahma-related gene-1, Blimp1, and Snail [33,34,35], and promotes symmetric dimethylation of arginine residues in histones (H3R8 and H4R3) [6]. PRMT5 was also shown to function as a transcriptional co-activator in MyoD-dependent regulation of Myog transcription in muscle tissues [7].
Recently, Tsai and colleague [36] reported that PRMT5 can function as a coactivator of CREB-dependent transcriptional events in the liver. PRMT5 was shown to specifically interact with CRTC2, a CREB coactivator that enhances CREB-dependent gluconeogenic programs under fasting conditions. Recruitment of PRMT5 to the promoters of PEPCK and G6Pase in turn promotes increased symmetric dimethylation of histone H3 at arginine 2 (H3R2), leading to increased lysine modification of the same histones by WD repeat-containing protein 5 (WDR5)-containing lysine methyl transferase complexes. Tsai and colleague [36] suggested that enhanced chromatin accessibility following H3R2 methylation is critical for stimulating CREB phosphorylation and subsequent transcription. Interestingly, a recent report suggested that PRMT5 can potentiate the repressor activity of small heterodimer partner (SHP), a known repressor of gluconeogenesis, by direct methylation of this protein at arginine 57 [37]. PRMT5 physically associates with SHP and promotes its methylation at Arg57. Methylated SHP in turn recruits transcriptional corepressors, including Brm, Sin3A, and HDAC1, leading to the inhibition of metabolic target genes. PRMT5-dependent regulation of SHP is essential for reducing bile acid and hepatic triglyceride levels and improving glucose tolerance in this model. Whether PRMT5-dependent regulation of CREB-dependent transcription can be compromised by PRMT5-mediated activation of SHP in animal models requires further study.
PRMT6
PRMT6 is a type I PRMT. Unlike other family members, PRMT6 was shown to contribute to transcriptional repression by enhancing asymmetric dimethylation of histone 3 at arginine 2 (H3R2), which in turn blocks WDR5-dependent H3K4 methylation and reduces transcription [38]. On the other hand, PRMT6 was also shown to promote asymmetric dimethylation of histone H4 at arginine 3 (H4R3), a marker for transcriptional activation, suggesting that it can also function as a coactivator of transcription [38]. Indeed, PRMT6 was shown to enhance the transcriptional activity of multiple nuclear hormone receptors in association with other coactivators including SRC-1 [39].
We recently showed that PRMT6 can also regulate hepatic gluconeogenesis via activation of a CRTC2-dependent transcriptional pathway in the liver [40]. PRMT6 was found to interact with CRTC2 in an endogenous coimmunoprecipitation experiment. We showed that PRMT6 directly modulates asymmetric dimethylation of multiple arginine residues in the N-terminus of CTRC2, thereby potentiating the interaction between CRTC2 and CREB on the promoters of gluconeogenic genes. Liver PRMT6 expression was increased in obese and insulin-resistant mice, and depletion of PRMT6 restored normal glycemia in these mouse models. This underscores the importance of PRMT6 in the control of CRTC2-dependent gluconeogenic transcription in vivo. We also observed similar modification of arginine residues in other CRTC proteins by PRMT6, suggesting that PRMT6-dependent regulation of CRTC activity can also affect other aspects of metabolism in cells such as fat cells or pancreatic β-cells.
CONCLUSIONS
In this review, we have summarized recent findings regarding the roles of PRMTs in the control of hepatic glucose metabolism. Interestingly, except for PRMT1, which regulates FoxO1, all of the PRMTs known to be involved in the control of hepatic glucose metabolism (PRMT4, PRMT5, and PRMT6) affect the transcriptional activity of CREB-CRTC2-dependent machinery (Fig. 1). It will be of great interest to delineate the relative contribution of each PRMT protein to the control of hepatic gluconeogenesis in vivo.
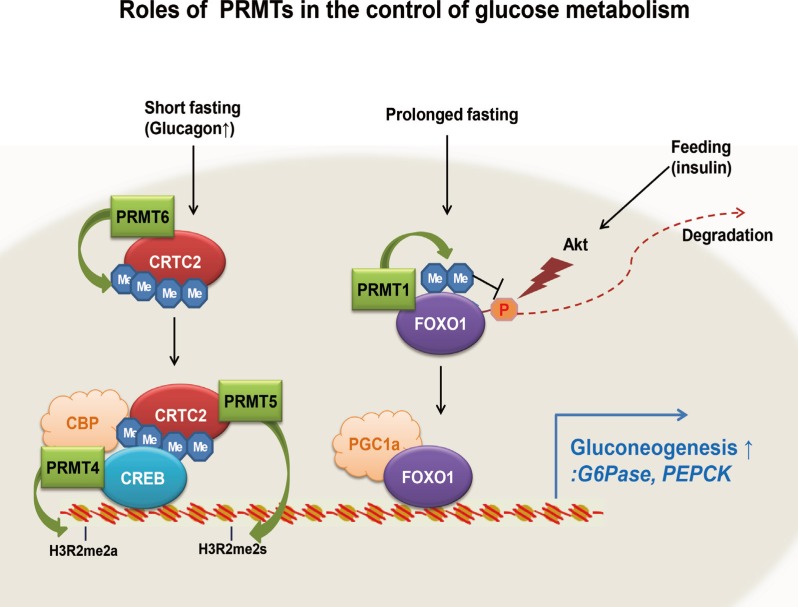
A model for the control of hepatic gluconeogenesis by protein arginine methyltransferases (PRMTs). The roles of PRMT1, PRMT4, PRMT5, and PRMT6 in controlling transcriptional activation of gluconeogenic genes and hepatic gluconeogenesis are shown. See the text for detailed information. CRTC2, CREB regulated transcription coactivator 2; FOXO1, forkhead box protein O1; CBP, CREB binding protein; CREB, cAMP response element-binding protein; PGC-1a, peroxisome proliferator-activated receptor γ coactivator 1α; G6Pase, glucose 6-phosphatase; PEPCK, phosphoenolpyruvate carboxykinase.
ACKNOWLEDGMENTS
This work was supported by the National Research Foundation of Korea (grant No. NRF-2010-0015098 and NRF-2012M3A9B6055345), funded by the Ministry of Science, ICT & Future Planning, Republic of Korea, and by grants from the Korean Health Technology R&D Project (grant No. A111345 and HI13C1886), funded by the Ministry of Health and Welfare, Republic of Korea.
Notes
No potential conflict of interest relevant to this article was reported.