Articles
- Page Path
- HOME > Endocrinol Metab > Volume 32(3); 2017 > Article
-
Original ArticleClinical Study Leu72Met and Other Intronic Polymorphisms in the GHRL and GHSR Genes Are Not Associated with Type 2 Diabetes Mellitus, Insulin Resistance, or Serum Ghrelin Levels in a Saudi Population
-
Faris Elbahi Joatar1*, Ali Ahmed Al Qarni1*, Muhalab E. Ali2, Abdulaziz Al Masaud1, Abdirashid M. Shire1, Nagalla Das3, Khalid Gumaa2,4, Hayder A. Giha2

-
Endocrinology and Metabolism 2017;32(3):360-369.
DOI: https://doi.org/10.3803/EnM.2017.32.3.360
Published online: September 18, 2017
1Department of Internal Medicine and Clinical Biochemistry Laboratory, King Abdulaziz Hospital, National Guard Health Affairs, Al-Ahsa, Saudi Arabia.
2Department of Medical Biochemistry, Arabian Gulf University College of Medicine and Medical Sciences, Manama, Bahrain.
3Clinical Biochemistry Laboratory, Salmaniya Medical Complex, Manama, Bahrain.
4Cell and Developmental Biology, Division of Biosciences, University College London, London, United Kingdom.
- Corresponding author: Hayder A. Giha. Department of Medical Biochemistry, Arabian Gulf University College of Medicine and Medical Sciences, PO Box 22979, Manama, Bahrain. Tel: +973-17239791, Fax: +973-17271090, gehaha2002@yahoo.com
- *These authors contributed equally to this work.
Copyright © 2017 Korean Endocrine Society
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
ABSTRACT
-
Background
- Ghrelin (GHRL), a gastric peptide encoded by the GHRL gene, is known to be involved in energy homeostasis via its G protein receptor, encoded by the growth hormone secretagogue receptor (GHSR) gene. Some studies have shown associations between plasma GHRL levels and GHRL single-nucleotide polymorphisms (SNPs), namely the Leu72Met polymorphism (rs696217 TG), with type 2 diabetes mellitus (T2DM) and insulin resistance (IR), while others have not. The controversies in these associations raise the issue of ‘which SNPs in which populations.’ The aim of this study was to investigate whether SNPs in GHRL and/or GHSR genes were associated with T2DM, IR, or plasma GHRL levels among Arab Saudis.
-
Methods
- Blood was collected from 208 Saudi subjects with (n=107) and without (n=101) T2DM. DNA samples from these subjects were analyzed by real-time polymerase chain reaction to genotype five intronic SNPs in the GHRL (rs696217 TG, rs27647 CT, rs2075356 CT, and rs4684677 AT) and GHSR (rs509030 GC) genes. In addition, plasma GHRL levels were measured by a radioimmunoassay.
-
Results
- None of the SNPs were associated with T2DM, IR, or plasma GHRL levels. The frequencies of the alleles, genotypes, and haplotypes of the five SNPs were comparable between the T2DM patients and the non-diabetic subjects. A large number of the GHRL haplotypes indicates the molecular heterogeneity of the preproghrelin gene in this region.
-
Conclusion
- Neither the Leu72Met polymorphism nor the other intronic GHRL and GHSR SNPs were associated with T2DM, IR, or GHRL levels. Further investigations should be carried out to explain the molecular basis of the association of the GHRL peptide with T2DM and IR.
- In 1999, ghrelin (GHRL) was identified as a peptide hormone secreted from stomach A-like cells and other tissues, and it was found that the effects of GHRL are mediated via growth hormone secretagogue receptors (GHSRs), which are located in the hypothalamus, heart, and adipose tissue [1]. GHRL was found to have an important direct role in energy homeostasis by affecting appetite and food intake [23] and indirectly by its influence on the actions of insulin, growth hormone, and probably other hormones. Thus, abnormalities of this peptide have pathological consequences that are reflected in type 2 diabetes mellitus (T2DM), adiposity, obesity, and body weight [23]. Additionally, GHRL was reported to be associated with hypertension [4]. A common clinical scenario for the above associations of GHRL is metabolic syndrome (MetS), in which the shared pathognomonic feature is insulin resistance (IR). However, GHRL has several other functions that were reviewed by Leite-Moreira and Soares in 2007 [5], including its role in cell proliferation and thus cancer risk. IR is a cardinal feature of T2DM and its associated metabolic abnormalities, the MetS, which together enhance cardiovascular morbidity, obesity, and mortality [67]. Although the exact mechanism of IR has yet to be confirmed, several risk factors have been identified, including GHRL deficiency.
- The preproghrelin (GHRL) gene is located on the short arm of chromosome 3 (3q25-26) and is transcribed into messenger RNA with four exons and five introns, encoding 117 amino acids, which are processed to give the 28-amino-acid molecule GHRL [18]. The active form is produced by addition of an octanoyl group (C7H15CO) to the third amino acid residue (serine). However, alternative promoters and alternative splicing of GHRL result in five transcript variants, some of which encode different protein isoforms; some do not encode any protein, but may regulate GHRL-obestatin preproprotein expression. Meanwhile, GHSRs, which are typical G protein-coupled receptors with seven transmembrane domains [9], are products of two transcriptional variants of the GHSR gene, on the short arm of chromosome 3 (3q25). The GHSR type 1a gene (GHSR), when an intron is excised, encodes the cognate receptor of GHRL [1]. The full-length receptor (type 1a) contains 366 amino acids encoded by two exons. The other transcriptional variant, type 1b, retains the intron; however, it may not function as a GHRL receptor.
- An association of low plasma GHRL levels with T2DM and IR was recently demonstrated among Saudi subjects [10]. Similar findings have been observed in other studies from different regions [111213]. Furthermore, few studies have investigated the relationship of GHRL and GHSR polymorphisms with plasma GHRL levels. Of those studies, several have reported associations of the Leu72Met (rs696217 TG) polymorphism with T2DM, IR, and other disorders [1415161718]. Thus, in this study, the aim was to examine the associations of Leu72Met and three other intronic GHRL single-nucleotide polymorphisms (SNPs) (rs27647 CT, rs2075356 CT, and rs4684677 AT) and one GHSR SNP (rs509030) with GHRL levels, T2DM, and IR among Saudi subjects. Knowledge of ethnic variations in the distribution and significance of the GHRL/GHSR SNPs will add to our knowledge about GHRL in general.
INTRODUCTION
- Study subjects
- This study was carried out in the Eastern Region of the Kingdom of Saudi Arabia, King Abdulaziz Hospital and primary health centers of the National Guard Health Affairs in Al-Ahsa Governorate. The 208 study subjects were aged between 18 and 75 years, and 107 subjects (61 males and 46 females) were known to have T2DM. The T2DM diagnosis was based on the 1998 World Health Organization diagnostic and classification criteria [19]. The patients were being followed up as outpatients. The other study subjects were 101 (57 males and 44 females) non-diabetic and apparently healthy individuals. Acute and chronic severe illnesses, pregnancy, and lactation were exclusion criteria.
- Study design and sample collection
- This was an observational cross-sectional study that received ethical approval from the research committees of two different institutions: the Arabian Gulf University and King Abdullah International Research Center, Riyadh, Kingdom of Saudi Arabia. After the aim of the study was explained, all patients and control subjects signed an informed consent form. Blood samples were drawn from all donors in the morning, after 10 to 12 hours of overnight fasting, into three tubes: (1) a gel/dry plain tube for the lipid profile; (2) a whole blood in ethylenediaminetetraacetic acid tube for hemoglobin A1c (HbA1c) and GHRL analysis, with the buffy coat retained for DNA extraction; and (3) a fluoride tube from which plasma was used to determine fasting glucose and insulin levels. Serum, plasma, and buffy coat samples were stored at −40℃ pending analysis.
- Blood chemistry
- Fasting blood glucose levels were estimated using an auto-analyzer, the Architect c8000 (Abbott Laboratories Inc., Abbott Park, IL, USA). HbA1c was estimated by measuring glycated hemoglobin using a G8 analyzer (TOSOH, Tessenderlo, Belgium).
- Plasma insulin was quantitatively assayed using a chemiluminescence procedure with an auto-analyzer, the Architect i2000 (Abbott Laboratories Inc.). IR was calculated using the homeostasis model assessment 2 (HOMA-IR) [20].
- Blood GHRL measurement
- The active form of plasma GHRL was measured by a radioimmunoassay (Millipore Research, St Charles, MO, USA). Briefly, on day 1, 100 µL of standards or plasma samples or controls and guinea pig GHRL antiserum were added to polystyrene tubes and incubated at 4℃ for 20 to 24 hours. On day 2, 100 µL of 125I-GHRL tracer was added and incubated at 4℃ for a further 20 to 24 hours. On day 3, 1 mL of cold precipitating reagent was added to each tube and incubated for 20 minutes at 4℃. After centrifugation at 2,000 to 3,000 rpm for 20 minutes at 4℃, radioactivity in the pellet was counted using a γ counter. The plasma GHRL levels (pmol/L) were then determined by the interpolation of the reference curve drawn using standard concentrations of GHRL. This method can achieve a sensitivity of 7.8 pg/mL using 100 µL plasma samples. This assay recognized only the active form of human GHRL [21] and showed no significant cross-reactivity with, or interference from, other factors related to GHRL (des-octanoyl GHRL <0.1%) or motilin-related peptide [4].
- Analysis of polymorphisms in the GHRL and GHSR genes
- Total genomic DNA was extracted from the leukocyte-rich buffy coat layer, using the Illustra protocol, limited DNA Mini Kit, according to the manufacturer's specifications (GE Healthcare UK Limited, Amersham Place, UK).
- The GHRL SNPs rs696217 TG (Leu72Met), rs27647 CT, rs2075356 CT (+3056), and rs3755777 GC (−1500) and the GHSR SNP rs509030 GC (Table 1) were selected by searching through the ABI HapMap (Applied Biosystems Inc., Foster City, CA, USA) data available via the SNP browser. The primer sequences for each SNP can be found at: http://www.ncbi.nlm.nih.gov/SNP/snp_ref.cgi?. All SNPs were in Hardy-Weinberg equilibrium (Table 2).
- GHRL/GHSR genotyping was done by real-time polymerase chain reaction (PCR), using the Applied Biosystems StepOne PCR System (Applied Biosystems Inc.). The assay is based on the detection of fluorescent signals generated proportionally during each PCR cycle. The genotyping was carried out in two steps: cycling (PCR amplification), followed by the endpoint detection of fluorescent signals. Allelic discrimination was achieved by selective annealing using TaqMan MGB probes (Applied Biosystems Inc.). Results were interpreted as homozygotes or heterozygotes [22].
- Statistical analysis
- The SigmaStat software version 3.5 (Systat Software Inc., San Jose, CA USA) was used for analysis. Comparisons were executed using the t test and one-way analysis of variance when the data were normally distributed; otherwise, the Mann-Whitney rank sum test and Kruskal-Wallis one-way analysis of variance on ranks were used. Rates and proportions were compared using the chi-square test. P values <0.05 were considered to indicate statistical significance.
- Hardy-Weinberg equilibrium values were calculated using an online calculator, OEGE (Online Encyclopedia for Genetic Epidemiology studies, http://www.oege.org/software/hwe-mr-calc.shtml), for the examined SNPs (the cut-off point for the probability of a population being in Hardy-Weinberg equilibrium was a chi-square value ≥3.841). The linkage equilibrium calculations and the identification of the haplotypes were done using the Haploview 4.2 program (Broad Institute, Cambridge, MA, USA).
- Ethical approval
- All procedures performed in studies involving human participants were conducted in accordance with the ethical standards of the relevant Institutional and National Research Committees and with the 1964 Helsinki Declaration and its later amendments.
METHODS
Blood glucose analysis
Plasma insulin
DNA extraction
Selection of the GHRL and GHSR SNPs
Real-time polymerase chain reaction analysis
- The frequency of GHRL and GHSR polymorphisms in T2DM patients and non-diabetic subjects
- As shown in Table 2, the minor allele frequency (MAF) of the examined GHRL (rs696217, rs2075356, rs27647, and rs-3755777) and GHSR (rs509030) intronic SNPs were not significantly different between subjects with and without T2DM (P=0.682, P=0.573, P=0.860, P=0.613, and P=0.825, respectively). Similarly, the distributions of the genotypes of the five examined SNPs were comparable between the study groups (P=0.237, P=0.987, P=0.879, P=0.505, and P=0.889, respectively).
- A total of 38 haplotypes were identified; 17 were found in both study groups, 10 were found only in the non-diabetic group, and 11 were found only in T2DM subjects. However, the proportions of the unique haplotypes were too small (each <3%) to be included in the comparisons. In contrast, the prevalence rates of the relatively more common haplotypes (≥3% in at least one study group) were compared between the two study groups and were found to be quite similar (Table 3).
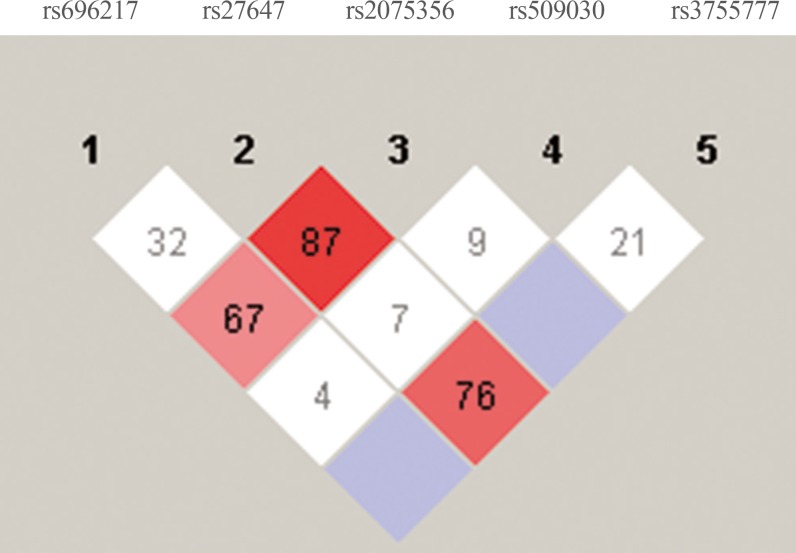
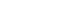
- As shown in Fig. 1, there was a strong linkage between rs27647 and rs2075356 (D′=87), and between rs27647 and rs3755777 (D′=76). There was also a linkage between rs696217 and rs2075356 (D′=67). Indeed, the smallest D′ values were those of rs509030 with the other SNPs, since this SNP is found in a different locus on chromosome 3 (3q25).
- Associations of the GHRL and GHSR SNPs with HbA1c
- For testing the associations of the GHRL and GHSR SNPs with HbA1c levels (as a countable marker of long-term blood glucose levels), all study subjects, regardless of whether they had T2DM, were analyzed together (Table 4). Among all study subjects, the HbA1c level was not significantly different between the carriers and non-carriers of the minor alleles of any of the five examined GHRL (rs2075356, rs696217, rs27647, and rs3755777) and GHSR (rs509030) SNPs (P=0.484, P=0.941, P=0.853, P=0.207, and P=0.488, respectively). Moreover, the genotypes (rare [minor] and common homozygous, and heterozygous) of the carriers of these SNPs had comparable HbA1c levels (P=0.539, P=not valid, P=0.784, P=0.202, and P=0.430, respectively).
- Associations of the GHRL and GHSR SNPs with IR (HOMA index)
- In all study subjects, the HOMA index, as a countable marker of IR, was not significantly different between the carriers and non-carriers of minor alleles of the GHRL (rs2075356, rs696217, rs27647, and rs3755777) and GHSR (rs509030) SNPs (P=0.132, P=0.517, P=0.165, P=0.434, and P=0.519, respectively). Furthermore, the HOMA index was comparable between the carriers of the rare (minor) and common homozygous, and heterozygous genotypes of the same SNPs (P=0.319, P=not valid, P=0.369, P=0.345, and P=0.386, respectively) (Table 5).
- Associations of the GHRL and GHSR SNPs with plasma GHRL levels
- In all study subjects, the plasma GHRL peptide levels (pmol/L) were not significantly different between the carriers and non-carriers of the minor alleles of the GHRL (rs2075356, rs696217, rs27647, and rs3755777) and GHSR (rs509030) SNPs (P=0.192, P=0.182, P=0.762, P=0.345, and P=0.808, respectively). Similarly, plasma GHRL levels were comparable between the GHRL and GHSR genotypes' carriers (rare and common homozygous, and heterozygous) of the tested SNPs (P=0.425, P=not valid, P=0.640, P=0.477, and P=0.971, respectively) (Table 6).
RESULTS
Alleles and genotypes
Haplotypes
Linkage disequilibrium between the study SNPs
- The exact mechanism of IR has not been precisely identified; however, clinical and laboratory correlates of IR have been reported [62324]. In a recent study, we demonstrated that IR and related markers were associated with low GHRL levels [10], as shown in other studies [4111213]. In this study, we found no associations between Leu72Met or other intronic GHRL/GHSR polymorphisms with T2DM, IR, or GHRL levels among Saudi subjects.
- We examined four SNPs (rs696217 [Leu72Met], rs2075356, rs27647, and rs3755777) in the GHRL gene and one in the GHSR gene (rs509030). All these polymorphisms are intronic in location, although the location of the rs509030 SNP is not precisely known. The selected SNPs were analyzed in samples that were previously used to document the associations of plasma GHRL levels with T2DM and IR. One of the aims of this study was to investigate whether the reported influence of GHRL in this population was affected by intronic SNPs of the GHRL gene, namely Leu72Met, and whether these SNPs were predictive of plasma GHRL levels. In several studies in different populations, the Leu72Met polymorphism was shown to be associated with T2DM, IR, and/or plasma GHRL levels [1415161718]. In this study, none of the five SNPs was associated with these disorders or plasma GHRL.
- Our findings are not unexpected, as there is no consensus that the examined SNPs are associated with T2DM or IR. The best example is Leu72Met, which has been thoroughly investigated before. The Leu72Met polymorphism was shown to be associated with T2DM [1415], IR [1617], and plasma GHRL levels [1618], and it was also found to be associated with diabetes-related disorders, such as MetS [1525], obesity [26], hypertension [14], and nephropathy [27]. Moreover, this SNP has been associated with diabetes-unrelated disorders, such as cancer [28], and bulimia nervosa purging type (BN-P) [29]. In contrast, several other studies showed that the Leu72Met polymorphism was not associated with T2DM [18303132], IR [14], or GHRL levels [433]. This SNP (Leu72Met) that has been widely tested across various nations had a MAF of only 0.04 in our study, which is more comparable to that found in Chinese (0.03) and notably different from the value found in Caucasians (0.25) and Japanese (0.20), as shown in the ABI HapMap data.
- Another commonly examined GHRL SNP (rs27647) was not associated with T2DM, IR, or plasma GHRL levels in this study. However, in other studies, it was found to be associated with plasma GHRL levels [16], IR [1634], hypertension [1435], and cancer [36]. In contrast, another study showed no association between this SNP and IR [14]. The lack of any previously reported associations of this SNP (rs27647) with T2DM was either because it was not tested for T2DM or because it was tested, but did not show a significant association, and thus was not reported. The rs27647 MAF among our study population (0.35) was similar to that of Caucasians (0.34, HapMap), and extremely different from that observed in African (0.140) and Japanese (0.11) subjects.
- The remaining two GHRL SNPs, rs2075356 and rs3755777, have been less commonly examined than the previous two; in this study, neither was found to be associated with T2DM, IR, or GHRL plasma levels. However, these two SNPs have been found to be associated with obesity [26]. Further, the rs2075356 polymorphism was also found to be associated with T2DM [14], BN-P [29], and cancer [37], while the rs3755777 polymorphism was not found to be associated with T2DM [38]. In this study, the MAFs of rs2075356 (0.10) and rs3755777 (0.17) were closer to the values observed among Caucasians (0.10 and 0.24, respectively), and notably different from the values observed among African-Americans (0.01 and 0.46, respectively) and Japanese (0.30 and 0.42, respectively).
- In contrast, GHSR polymorphisms have been investigated less often. In this study, we tested the rs509030 GC polymorphism. This SNP has rarely been tested, and we found only one peer-reviewed published article on this SNP in PubMed [39]. In our study, the rs509030 GC polymorphism had the least amount of linkage with the other SNPs (Fig. 1), which might indicate that it is not located on chromosome 3. Furthermore, it was not associated with T2DM or IR, and it did not predict plasma GHRL levels. However, the rs509030 GC polymorphism was found to be associated with MetS and dyslipidemia [39]. Although the MAF of this SNP was 0.31, which is relatively high compared to the other tested SNPs in this study (Table 2), there were no MAF data available for this SNP online.
- Of the 38 identified GHRL/GHSR haplotypes, the prevalence of 11 common haplotypes, as well as the remaining haplotypes, were comparable between the T2DM and non-T2DM subjects. A large number of haplotypes indicates the genetic diversity of the preproghrelin gene, although this diversity does not affect the structure or function of GHRL (comparable between study subjects) or the protein expression (plasma GHRL levels) of this gene. However, more polymorphisms need to be investigated and/or identified in order to explain the significant differences observed in GHRL levels between T2DM subjects with and without IR and non-diabetic subjects. Furthermore, none of the alleles/genotypes of the tested SNPs were associated with the most important measurable parameter of T2DM, HbA1c (Table 4), or with that of IR, the HOMA index (Table 5).
- Finally, it remains necessary to find an explanation for the inconsistent associations between the GHRL/GHSR polymorphisms, especially the Leu72Met polymorphism, and T2DM and IR. A recent meta-analysis study of the Leu72Met polymorphism and T2DM used six case-control studies (a total of 3,417 cases and 3,081 controls), and demonstrated that no association was present between the Leu72Met polymorphism and T2DM in the overall population or in Asians. However, in subgroup analyses stratified by ethnicity, they found that the risk for T2DM was decreased in subjects with Met72+ genotypes in Caucasians [40]. Indeed, the differences between the studies' subjects, whether genetic (ethnicity), due to environmental factors, or even the way the study subjects were selected, could in part explain the controversies in the literature.
- In conclusion, this study documented an important negative finding: the lack of an association of T2DM in Saudis with Leu72Met, the most investigated GHRL SNP, which was previously found to be associated with T2DM and IR in some populations but not the others. It also affirmed the role of ethnicity in the interpretation of SNP association studies, and investigated a rarely analyzed GHSR SNP, rs509030 GC; however, this SNP and other GHRL intronic SNPs (rs2075356, rs27647, and rs3755777) were likewise not associated with T2DM, IR, or plasma GHRL, while their haplotypes indicated great genetic variability. The limitations of this study include the small sample size and the limited number of tested SNPs. Thus, a further analysis of more GHRL-related SNPs with a larger sample size is needed to explain the difference in the plasma GHRL peptide levels seen in our study subjects.
DISCUSSION
-
Acknowledgements
- We acknowledge the contributions of our study subjects, both diabetic and healthy volunteers. We are also grateful for the clinicians from the National Guard Health Affairs, Primary Health Care, and the departmental staff in King Abdulaziz National Guard Hospital, Al-Ahsa, KSA, who assisted in data collection and biochemical analysis. We must extend our acknowledgements to Mrs. Lamees J Habieb and Mr. Abdullah J Hassan at Salmaniya Medical Complex (SMC), Kingdom of Bahrain, for their technical assistance. Finally, we are indebted to our study sponsors from Arabian Gulf University, Bahrain (AGU, grant 74-2011), King Abdullah International Medical Research Center, Saudi Arabia (grant number, IRBC-112-11), and WHO/EMRO (grant number TSA 200303689).
ACKNOWLEDGMENTS
-
CONFLICTS OF INTEREST: No potential conflict of interest relevant to this article was reported.
Article information
- 1. Kojima M, Hosoda H, Date Y, Nakazato M, Matsuo H, Kangawa K. Ghrelin is a growth-hormone-releasing acylated peptide from stomach. Nature 1999;402:656–660. ArticlePubMedPDF
- 2. Tschop M, Smiley DL, Heiman ML. Ghrelin induces adiposity in rodents. Nature 2000;407:908–913. ArticlePubMedPDF
- 3. Lopez M, Lage R, Saha AK, Perez-Tilve D, Vazquez MJ, Varela L, et al. Hypothalamic fatty acid metabolism mediates the orexigenic action of ghrelin. Cell Metab 2008;7:389–399. ArticlePubMed
- 4. Poykko SM, Kellokoski E, Horkko S, Kauma H, Kesaniemi YA, Ukkola O. Low plasma ghrelin is associated with insulin resistance, hypertension, and the prevalence of type 2 diabetes. Diabetes 2003;52:2546–2553. ArticlePubMed
- 5. Leite-Moreira AF, Soares JB. Physiological, pathological and potential therapeutic roles of ghrelin. Drug Discov Today 2007;12:276–288. ArticlePubMed
- 6. Grundy SM. Atherogenic dyslipidemia associated with metabolic syndrome and insulin resistance. Clin Cornerstone 2006;8(Suppl 1):S21–S27. ArticlePubMed
- 7. Grundy SM. Metabolic syndrome: connecting and reconciling cardiovascular and diabetes worlds. J Am Coll Cardiol 2006;47:1093–1100. ArticlePubMed
- 8. Wajnrajch MP, Ten IS, Gertner JM, Leibel RL. Genomic organization of the human GHRELIN gene. J Endocr Genet 2000;1:231–233.ArticlePDF
- 9. Davenport AP, Bonner TI, Foord SM, Harmar AJ, Neubig RR, Pin JP, et al. International Union of Pharmacology. LVI. Ghrelin receptor nomenclature, distribution, and function. Pharmacol Rev 2005;57:541–546. ArticlePubMed
- 10. Al Qarni AA, Joatar FE, Das N, Awad M, Eltayeb M, Al-Zubair AG, et al. Association of plasma ghrelin levels with insulin resistance in type 2 diabetes mellitus among Saudi subjects. Endocrinol Metab (Seoul) 2017;32:230–240. ArticlePubMedPMC
- 11. Dezaki K, Sone H, Yada T. Ghrelin is a physiological regulator of insulin release in pancreatic islets and glucose homeostasis. Pharmacol Ther 2008;118:239–249. ArticlePubMed
- 12. Yada T, Dezaki K, Sone H, Koizumi M, Damdindorj B, Nakata M, et al. Ghrelin regulates insulin release and glycemia: physiological role and therapeutic potential. Curr Diabetes Rev 2008;4:18–23. ArticlePubMed
- 13. Pacifico L, Poggiogalle E, Costantino F, Anania C, Ferraro F, Chiarelli F, et al. Acylated and nonacylated ghrelin levels and their associations with insulin resistance in obese and normal weight children with metabolic syndrome. Eur J Endocrinol 2009;161:861–870. ArticlePubMed
- 14. Mager U, Kolehmainen M, Lindstrom J, Eriksson JG, Valle TT, Hamalainen H, et al. Association between ghrelin gene variations and blood pressure in subjects with impaired glucose tolerance. Am J Hypertens 2006;19:920–926. ArticlePubMedPDF
- 15. Takezawa J, Yamada K, Morita A, Aiba N, Watanabe S. Preproghrelin gene polymorphisms in obese Japanese: association with diabetes mellitus in men and with metabolic syndrome parameters in women. Obes Res Clin Pract 2009;3:179–191. ArticlePubMed
- 16. Zavarella S, Petrone A, Zampetti S, Gueorguiev M, Spoletini M, Mein CA, et al. A new variation in the promoter region, the -604 C>T, and the Leu72Met polymorphism of the ghrelin gene are associated with protection to insulin resistance. Int J Obes (Lond) 2008;32:663–668. ArticlePubMedPDF
- 17. Liu J, Liu J, Tian LM, Liu JX, Bing YJ, Zhang JP, et al. Association of ghrelin Leu72Met polymorphism with type 2 diabetes mellitus in Chinese population. Gene 2012;504:309–312. ArticlePubMed
- 18. Hedayatizadeh-Omran A, Rafiei A, Khajavi R, Alizadeh-Navaei R, Mokhberi V, Moradzadeh K. Association between ghrelin gene (Leu72Met) polymorphism and ghrelin serum level with coronary artery diseases. DNA Cell Biol 2014;33:95–101. ArticlePubMed
- 19. Alberti KG, Zimmet PZ. Definition, diagnosis and classification of diabetes mellitus and its complications. Part 1: diagnosis and classification of diabetes mellitus provisional report of a WHO consultation. Diabet Med 1998;15:539–553. ArticlePubMed
- 20. Matthews DR, Hosker JP, Rudenski AS, Naylor BA, Treacher DF, Turner RC. Homeostasis model assessment: insulin resistance and beta-cell function from fasting plasma glucose and insulin concentrations in man. Diabetologia 1985;28:412–419. ArticlePubMedPDF
- 21. Prudom C, Liu J, Patrie J, Gaylinn BD, Foster-Schubert KE, Cummings DE, et al. Comparison of competitive radioimmunoassays and two-site sandwich assays for the measurement and interpretation of plasma ghrelin levels. J Clin Endocrinol Metab 2010;95:2351–2358. ArticlePubMedPMCPDF
- 22. Finan RR, Mustafa FE, Al-Zaman I, Madan S, Issa AA, Almawi WY. STAT3 polymorphisms linked with idiopathic recurrent miscarriages. Am J Reprod Immunol 2010;63:22–27. ArticlePubMed
- 23. Weyer C, Bogardus C, Mott DM, Pratley RE. The natural history of insulin secretory dysfunction and insulin resistance in the pathogenesis of type 2 diabetes mellitus. J Clin Invest 1999;104:787–794. ArticlePubMedPMC
- 24. Ma ZA, Zhao Z, Turk J. Mitochondrial dysfunction and β-cell failure in type 2 diabetes mellitus. Exp Diabetes Res 2012;2012:703538. ArticlePubMedPDF
- 25. Steinle NI, Pollin TI, O’Connell JR, Mitchell BD, Shuldiner AR. Variants in the ghrelin gene are associated with metabolic syndrome in the Old Order Amish. J Clin Endocrinol Metab 2005;90:6672–6677. ArticlePubMed
- 26. Takezawa J, Yamada K, Miyachi M, Morita A, Aiba N, Sasaki S, et al. Preproghrelin gene polymorphisms in obese Japanese women. Minor homozygotes are light eaters, do not prefer protein or fat, and apparently have a poor appetite. Appetite 2013;63:105–111. ArticlePubMed
- 27. Zhuang L, Li M, Yu C, Li C, Zhao M, Lu M, et al. The Leu72Met polymorphism of the GHRL gene prevents the development of diabetic nephropathy in Chinese patients with type 2 diabetes mellitus. Mol Cell Biochem 2014;387:19–25. ArticlePubMedPDF
- 28. Wu IC, Zhao Y, Zhai R, Liu G, Ter-Minassian M, Asomaning K, et al. Association between polymorphisms in cancer-related genes and early onset of esophageal adenocarcinoma. Neoplasia 2011;13:386–392. ArticlePubMedPMC
- 29. Ando T, Komaki G, Naruo T, Okabe K, Takii M, Kawai K, et al. Possible role of preproghrelin gene polymorphisms in susceptibility to bulimia nervosa. Am J Med Genet B Neuropsychiatr Genet 2006;141B:929–934. ArticlePubMed
- 30. Larsen LH, Gjesing AP, Sorensen TI, Hamid YH, Echwald SM, Toubro S, et al. Mutation analysis of the preproghrelin gene: no association with obesity and type 2 diabetes. Clin Biochem 2005;38:420–424. ArticlePubMed
- 31. Kim SY, Jo DS, Hwang PH, Park JH, Park SK, Yi HK, et al. Preproghrelin Leu72Met polymorphism is not associated with type 2 diabetes mellitus. Metabolism 2006;55:366–370. ArticlePubMed
- 32. Garcia EA, King P, Sidhu K, Ohgusu H, Walley A, Lecoeur C, et al. The role of ghrelin and ghrelin-receptor gene variants and promoter activity in type 2 diabetes. Eur J Endocrinol 2009;161:307–315. ArticlePubMed
- 33. Zou CC, Huang K, Liang L, Zhao ZY. Polymorphisms of the ghrelin/obestatin gene and ghrelin levels in Chinese children with short stature. Clin Endocrinol (Oxf) 2008;69:99–104. ArticlePubMed
- 34. Gueorguiev M, Lecoeur C, Meyre D, Benzinou M, Mein CA, Hinney A, et al. Association studies on ghrelin and ghrelin receptor gene polymorphisms with obesity. Obesity (Silver Spring) 2009;17:745–754. ArticlePubMed
- 35. Berthold HK, Giannakidou E, Krone W, Tregouet DA, Gouni-Berthold I. Influence of ghrelin gene polymorphisms on hypertension and atherosclerotic disease. Hypertens Res 2010;33:155–160. ArticlePubMedPDF
- 36. Campa D, Pardini B, Naccarati A, Vodickova L, Novotny J, Steinke V, et al. Polymorphisms of genes coding for ghrelin and its receptor in relation to colorectal cancer risk: a two-step gene-wide case-control study. BMC Gastroenterol 2010;10:112ArticlePubMedPMCPDF
- 37. Pabalan NA, Seim I, Jarjanazi H, Chopin LK. Associations between ghrelin and ghrelin receptor polymorphisms and cancer in Caucasian populations: a meta-analysis. BMC Genet 2014;15:118ArticlePubMedPMCPDF
- 38. Choi HJ, Cho YM, Moon MK, Choi HH, Shin HD, Jang HC, et al. Polymorphisms in the ghrelin gene are associated with serum high-density lipoprotein cholesterol level and not with type 2 diabetes mellitus in Koreans. J Clin Endocrinol Metab 2006;91:4657–4663. ArticlePubMedPDF
- 39. Li WJ, Zhen YS, Sun K, Xue H, Song XD, Wang YB, et al. Ghrelin receptor gene polymorphisms are associated with female metabolic syndrome in Chinese population. Chin Med J (Engl) 2008;121:1666–1669. ArticlePubMed
- 40. Liao N, Xie ZK, Huang J, Xie ZF. Association between the ghrelin Leu72Met polymorphism and type 2 diabetes risk: a meta-analysis. Gene 2013;517:179–183. ArticlePubMed
References
Linkage disequilibrium analyses of ghrelin single-nucleotide polymorphisms (SNPs; rs2075356, rs696217, rs27647, rs3755777) and a growth hormone secretagogue receptor (GHSR) SNP (rs509030 GC). The strength of the linkage is indicated by the value of D′, and the maximum value (absolute linkage) is 100%. The GHSR SNP had the least linkage with the other ghrelin SNPs.
The Selected Ghrelin and Growth Hormone Secretagogue Receptor SNPs: Chromosome, Band, and Locus and Polymorphism Type and Class
Hardy-Weinberg Equilibrium, MAF, and Prevalence of Rare Genotypes of the Ghrelin and Growth Hormone Secretagogue Receptor SNPs among T2DM and Non-diabetic Subjects
The Prevalence of Ghrelin and Growth Hormone Secretagogue Receptor SNPs Haplotypes in T2DM and Non-diabetic Subjects
Associations of Ghrelin and Growth Hormone Secretagogue Receptor Polymorphisms with HbA1c: Comparisons between Carriers and Non-Carriers of Minor Alleles and Rare Genotypes of the Examined SNPs
Values are expressed as median (interquartile range).
HbA1c, hemoglobin A1c; SNP, single-nucleotide polymorphism.
aMann-Whitney rank sum test; bKruskal-Wallis 1-way analysis of variance on ranks; cThe numbers of carriers of rare homozygous, heterozygous, and common homozygous genotypes, respectively, for each SNP.
Associations of Ghrelin and Growth Hormone Secretagogue Receptor Polymorphisms with Insulin Resistance (HOMA): Comparisons between Carriers and Non-Carriers of Minor Alleles and between the Genotypes of the Carriers of the Examined SNPs
Values are expressed as median (interquartile range).
HOMA, homeostatic model assessment; SNP, single-nucleotide polymorphism.
aMann-Whitney rank sum test; bKruskal-Wallis 1-way analysis of variance on ranks; cThe numbers of carriers of the rare homozygous, heterozygous, and common homozygous genotypes, respectively, for each SNP.
Associations of Ghrelin and Growth Hormone Secretagogue Receptor Polymorphisms with Plasma Ghrelin Levels: Comparisons between Carriers and Non-Carriers of Minor Alleles and between the Genotypes of the Carriers of the Examined SNPs
Figure & Data
References
Citations

- Relationship between single nucleotide polymorphism studies in ghrelin gene with obesity subjects
May Salem Al-Nbaheen
Journal of King Saud University - Science.2023; 35(1): 102393. CrossRef - Leu72Met Polymorphism in Ghrelin Gene: A Potential Risk Factor for Hypertension in Type 2 Diabetes Patients
Monika Buraczynska, Jakub Golacki, Wojciech Zaluska
Diabetes, Metabolic Syndrome and Obesity.2023; Volume 16: 557. CrossRef - Асоціації варіантів гена GHRL із розвитком ожиріння та метаболічних порушень у дітей
A. Abaturov, A. Nikulina
CHILD`S HEALTH.2023; 18(4): 255. CrossRef - Effect of the GHRL gene (rs696217) polymorphism on the metabolic disorders in patients with obesity in the Ukrainian population
Andrii Prodan, Ihor Dzubanovsky, Oleksandr Kamyshnyi, Natalia Melnyk, Stepan Grytsenko, Stanislava Voloshyn
Endocrine Regulations.2023; 57(1): 173. CrossRef - Impact of gene polymorphism of glutathione S-transferase and ghrelin as a risk factor in Egyptian women with gestational diabetes mellitus
Mai M. Madkour, Afaf M. El-Said, Abd El-Aziz A. El-Refaey, Abd El-Aziz F. Abd El-Aziz, Fardous F. El-Senduny
Egyptian Journal of Medical Human Genetics.2022;[Epub] CrossRef -
Association of obesity in T2DM with differential polymorphism of ghrelin, growth hormone secretagogue receptor-1 and telomeres maintenance genes
Hayder A. Giha, Faris E. Joatar, Dhuha M. B. AlDehaini, Zainab H. A. Malalla, Muhalab E. Ali, Ali A. Al Qarni
Hormone Molecular Biology and Clinical Investigation.2022; 43(3): 297. CrossRef - Investigation of GHRL (rs4684677), FTO (rs8044769) and PGC1Α (rs8192678) polymorphisms in type 2 diabetic Turkish population
Osman Oğuz, Arezoo Gheybi, Zeliha Doğan, Feray Akbaş, Ümit Zeybek, Arzu Ergen
Turkish Journal of Biochemistry.2022; 47(5): 564. CrossRef - Grelin ve Grelin Reseptörü Polimorfizmlerinin Tip 2 Diyabetle İlişkisi
Esma SELÇUK, Uğur ŞAHİN, Didem ÖZKAHRAMAN, Mustafa CALAPOĞLU, Nilüfer ŞAHİN CALAPOĞLU
Süleyman Demirel Üniversitesi Sağlık Bilimleri Dergisi.2022; 13(2): 218. CrossRef - Variation analysis of Ghrelin gene in Chinese patients with obesity, having polycystic ovarian syndrome
Xiaomeng Wang, Fengxiang Qu, Chunlian Wang, Yan Wang, Dan Wang, Min Zhao, Xiangbing Yun, Qingmei Zheng, Lin Xu
Gynecological Endocrinology.2020; 36(7): 594. CrossRef - The flavonoid baicalin improves glucose metabolism by targeting the PH domain of AKT and activating AKT/GSK3β phosphorylation
Shengnan Yang, Yuan Zhang, Fukui Shen, Xiaoyao Ma, Man Zhang, Yuanyuan Hou, Gang Bai
FEBS Letters.2019; 593(2): 175. CrossRef - Ethnicity-Specific Association Between Ghrelin Leu72Met Polymorphism and Type 2 Diabetes Mellitus Susceptibility: An Updated Meta-Analysis
Rong Huang, Sai Tian, Rongrong Cai, Jie Sun, Yanjue Shen, Shaohua Wang
Frontiers in Genetics.2018;[Epub] CrossRef

 KES
KES
 PubReader
PubReader Cite
Cite